|
Triple-stranded DNA
Triple-stranded DNA (also known as H-DNA or Triplex-DNA) is a DNA structure in which three oligonucleotides wind around each other and form a triple helix. In triple-stranded DNA, the third strand binds to a B-form DNA (via Watson–Crick base-pairing) double helix by forming Hoogsteen base pairs or reversed Hoogsteen hydrogen bonds. Structure Examples of triple-stranded DNA have been found in natural sources with the required structural elements, for example in Satellite DNA. Hoogsteen base pairing A thymine (T) nucleobase can bind to a Watson–Crick base-pairing of T-A by forming a Hoogsteen hydrogen bond. The thymine hydrogen bonds with the adenosine (A) of the original double-stranded DNA to create a T-A*T base-triplet. Intermolecular and intramolecular interactions There are two classes of triplex DNA: intermolecular and intramolecular formations. An intermolecular triplex refers to triplex formation between a duplex and a different (third) strand of DNA. Th ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Intron
An intron is any nucleotide sequence within a gene that is not expressed or operative in the final RNA product. The word ''intron'' is derived from the term ''intragenic region'', i.e. a region inside a gene."The notion of the cistron .e., gene... must be replaced by that of a transcription unit containing regions which will be lost from the mature messenger – which I suggest we call introns (for intragenic regions) – alternating with regions which will be expressed – exons." (Gilbert 1978) The term ''intron'' refers to both the DNA sequence within a gene and the corresponding RNA sequence in RNA transcripts. The non-intron sequences that become joined by this RNA processing to form the mature RNA are called exons. Introns are found in the genes of most organisms and many viruses and they can be located in both protein-coding genes and genes that function as RNA (noncoding genes). There are four main types of introns: tRNA introns, group I introns, group II introns, and ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Bcl-2
Bcl-2 (B-cell lymphoma 2), encoded in humans by the ''BCL2'' gene, is the founding member of the Bcl-2 family of regulator proteins that regulate cell death (apoptosis), by either inhibiting (anti-apoptotic) or inducing (pro-apoptotic) apoptosis. It was the first apoptosis regulator identified in any organism. Bcl-2 derives its name from ''B-cell lymphoma 2'', as it is the second member of a range of proteins initially described in chromosomal translocations involving chromosomes 14 and 18 in follicular lymphomas. Orthologs (such as ''Bcl2'' in mice) have been identified in numerous mammals for which complete genome data are available. Like BCL3, BCL5, BCL6, BCL7A, BCL9, and BCL10, it has clinical significance in lymphoma. Isoforms The two isoforms of Bcl-2, Isoform 1, and Isoform 2, exhibit a similar fold. However, results in the ability of these isoforms to bind to the BAD and BAK proteins, as well as in the structural topology and electrostatic potential of the binding ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Z-DNA
Z-DNA is one of the many possible double helical structures of DNA. It is a left-handed double helical structure in which the helix winds to the left in a zigzag pattern, instead of to the right, like the more common B-DNA form. Z-DNA is thought to be one of three biologically active double-helical structures along with A-DNA and B-DNA. History Left-handed DNA was first discovered by Robert Wells and colleagues, during their studies of a repeating polymer of inosine–cytosine. They observed a "reverse" circular dichroism spectrum for such DNAs, and interpreted this (correctly) to mean that the strands wrapped around one another in a left-handed fashion. The relationship between Z-DNA and the more familiar B-DNA was indicated by the work of Pohl and Jovin, who showed that the ultraviolet circular dichroism of poly(dG-dC) was nearly inverted in 4 M sodium chloride solution. The suspicion that this was the result of a conversion from B-DNA to Z-DNA was confirmed by examini ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RecA
RecA is a 38 kilodalton protein essential for the repair and maintenance of DNA. A RecA structural and functional homolog has been found in every species in which one has been seriously sought and serves as an archetype for this class of homologous DNA repair proteins. The homologous protein is called RAD51 in eukaryotes and RadA in archaea. RecA has multiple activities, all related to DNA repair. In the bacterial SOS response, it has a co-protease function in the autocatalytic cleavage of the LexA repressor and the λ repressor. RecA's association with DNA repair is based on its central role in homologous recombination. The RecA protein binds strongly and in long clusters to ssDNA to form a nucleoprotein filament. The protein has more than one DNA binding site, and thus can hold a single strand and double strand together. This feature makes it possible to catalyze a DNA synapsis reaction between a DNA double helix and a complementary region of single-stranded DNA. The RecA-s ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Homologous Recombination
Homologous recombination is a type of genetic recombination in which genetic information is exchanged between two similar or identical molecules of double-stranded or single-stranded nucleic acids (usually DNA as in cellular organisms but may be also RNA in viruses). Homologous recombination is widely used by cells to accurately DNA repair harmful breaks that occur on both strands of DNA, known as double-strand breaks (DSB), in a process called homologous recombinational repair (HRR). Homologous recombination also produces new combinations of DNA sequences during meiosis, the process by which eukaryotes make gamete cells, like sperm and egg cells in animals. These new combinations of DNA represent genetic variation in offspring, which in turn enables populations to adapt during the course of evolution. Homologous recombination is also used in horizontal gene transfer to exchange genetic material between different strains and species of bacteria and viruses. Horizontal ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Wiki Image RecA
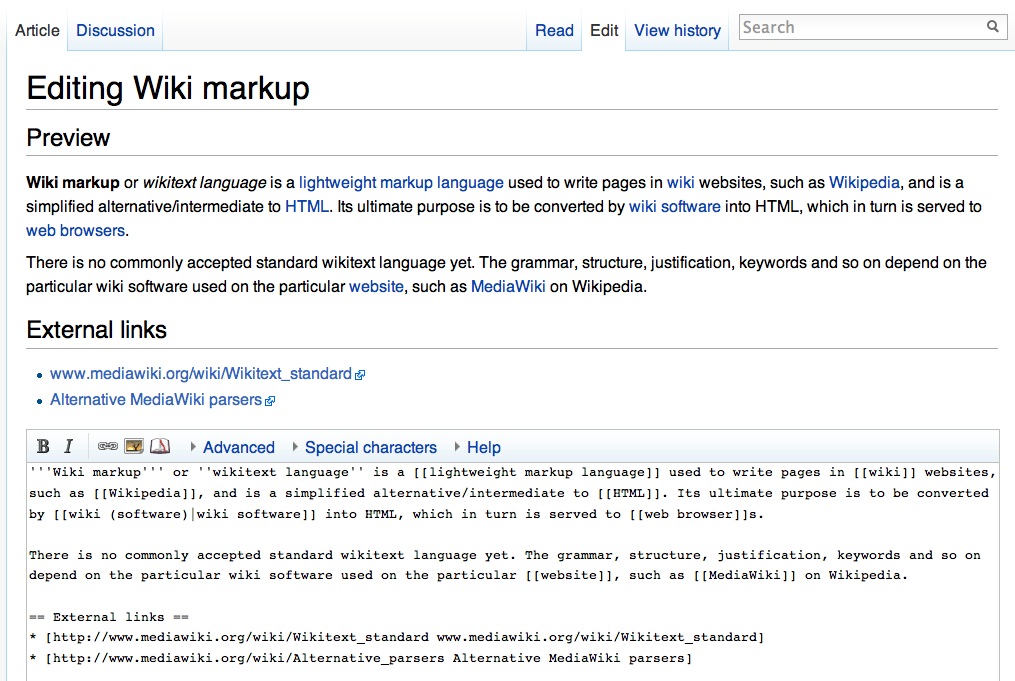
A wiki ( ) is an online hypertext publication collaboratively edited and managed by its own audience, using a web browser. A typical wiki contains multiple pages for the subjects or scope of the project, and could be either open to the public or limited to use within an organization for maintaining its internal knowledge base. Wikis are enabled by wiki software, otherwise known as wiki engines. A wiki engine, being a form of a content management system, differs from other web-based systems such as blog software, in that the content is created without any defined owner or leader, and wikis have little inherent structure, allowing structure to emerge according to the needs of the users. Wiki engines usually allow content to be written using a simplified markup language and sometimes edited with the help of a rich-text editor. There are dozens of different wiki engines in use, both standalone and part of other software, such as bug tracking systems. Some wiki engines are ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Na+/K+-ATPase
NA, N.A., Na, nA or n/a may refer to: Chemistry and physics * Sodium, symbol Na, a chemical element * Avogadro constant (''N''A) * Nucleophilic addition, a type of reaction in organic chemistry * Numerical aperture, a number that characterizes a range of angles in an optical system * nA, the symbol for nanoampere * Naturally aspirated engine Biology and medicine * Na (tree) or ''Mesua ferrea'', a species of tree native to Sri Lanka * Neuroacanthocytosis, a neurological condition * ''Nomina Anatomica'', a former international standard for human anatomical nomenclature * Noradrenaline, a hormone * Nucleic acid analogue, compounds analogous to naturally occurring RNA and DNA Places Current * Namibia (ISO country code) * Naples (car number plate code: NA), Italy * North America, a continent * North Africa, a subcontinent Historical * Netherlands Antilles (former international vehicle registration code: NA) * Na (Chinese state), a small state of the Chinese Zhou dynasty from ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Beta-lactamase
Beta-lactamases, (β-lactamases) are enzymes () produced by bacteria that provide multi-resistance to beta-lactam antibiotics such as penicillins, cephalosporins, cephamycins, monobactams and carbapenems (ertapenem), although carbapenems are relatively resistant to beta-lactamase. Beta-lactamase provides antibiotic resistance by breaking the antibiotics' structure. These antibiotics all have a common element in their molecular structure: a four-atom ring known as a beta-lactam (β-lactam) ring. Through hydrolysis, the enzyme lactamase breaks the β-lactam ring open, deactivating the molecule's antibacterial properties. Beta-lactam antibiotics are typically used to target a broad spectrum of gram-positive and gram-negative bacteria. Beta-lactamases produced by gram-negative bacteria are usually secreted, especially when antibiotics are present in the environment. Structure The structure of a '' Streptomyces'' serine β-lactamase (SBLs) is given by . The alpha-beta fold ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Lac Operon
The ''lactose'' operon (''lac'' operon) is an operon required for the transport and metabolism of lactose in ''E. coli'' and many other enteric bacteria. Although glucose is the preferred carbon source for most bacteria, the ''lac'' operon allows for the effective digestion of lactose when glucose is not available through the activity of beta-galactosidase. Gene regulation of the ''lac'' operon was the first genetic regulatory mechanism to be understood clearly, so it has become a foremost example of prokaryotic gene regulation. It is often discussed in introductory molecular and cellular biology classes for this reason. This lactose metabolism system was used by François Jacob and Jacques Monod to determine how a biological cell knows which enzyme to synthesize. Their work on the ''lac'' operon won them the Nobel Prize in Physiology in 1965. Bacterial operons are polycistronic transcripts that are able to produce multiple proteins from one mRNA transcript. In this case, when l ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Saccharomyces Cerevisiae
''Saccharomyces cerevisiae'' () (brewer's yeast or baker's yeast) is a species of yeast (single-celled fungus microorganisms). The species has been instrumental in winemaking, baking, and brewing since ancient times. It is believed to have been originally isolated from the skin of grapes. It is one of the most intensively studied eukaryotic model organisms in molecular biology, molecular and cell biology, much like ''Escherichia coli'' as the model bacteria, bacterium. It is the microorganism behind the most common type of fermentation (biochemistry), fermentation. ''S. cerevisiae'' cells are round to ovoid, 5–10 micrometre, μm in diameter. It reproduces by budding. Many proteins important in human biology were first discovered by studying their Homology (biology), homologs in yeast; these proteins include cell cycle proteins, signaling proteins, and protein-processing enzymes. ''S. cerevisiae'' is currently the only yeast cell known to have Berkeley body, Berkeley bo ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Promoter (genetics)
In genetics, a promoter is a sequence of DNA to which proteins bind to initiate transcription of a single RNA transcript from the DNA downstream of the promoter. The RNA transcript may encode a protein (mRNA), or can have a function in and of itself, such as tRNA or rRNA. Promoters are located near the transcription start sites of genes, upstream on the DNA (towards the 5' region of the sense strand). Promoters can be about 100–1000 base pairs long, the sequence of which is highly dependent on the gene and product of transcription, type or class of RNA polymerase recruited to the site, and species of organism. Promoters control gene expression in bacteria and eukaryotes. RNA polymerase must attach to DNA near a gene for transcription to occur. Promoter DNA sequences provide an enzyme binding site. The -10 sequence is TATAAT. -35 sequences are conserved on average, but not in most promoters. Artificial promoters with conserved -10 and -35 elements transcribe more slowly. All D ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
.png)