|
SIR Protein
Silent Information Regulator (SIR) proteins are involved in regulating gene expression. SIR proteins organize heterochromatin near telomeres, rDNA, and at silent loci including hidden mating type loci in yeast. The SIR family of genes encodes catalytic and non-catalytic proteins that are involved in de-acetylation of histone tails and the subsequent condensation of chromatin around a SIR protein scaffold. Some SIR family members are conserved from yeast to humans. History SIR proteins have been identified in many screens, and have historically been known as SIR (silent information regulator), MAR (mating-type regulator), STE (sterile), CMT (change of mating type) or SSP (sterile suppressor) according to which screen led to their identification. Ultimately, the name SIR had the most staying power, because it most accurately describes the function of the encoded proteins. One of the early yeast screens to identify SIR genes was performed by Anita Hopper and Benjamin Hall, who ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Heterochromatin
Heterochromatin is a tightly packed form of DNA or '' condensed DNA'', which comes in multiple varieties. These varieties lie on a continue between the two extremes of constitutive heterochromatin and facultative heterochromatin. Both play a role in the expression of genes. Because it is tightly packed, it was thought to be inaccessible to polymerases and therefore not transcribed; however, according to Volpe et al. (2002), and many other papers since, much of this DNA is in fact transcribed, but it is continuously turned over via RNA-induced transcriptional silencing (RITS). Recent studies with electron microscopy and OsO4 staining reveal that the dense packing is not due to the chromatin. Constitutive heterochromatin can affect the genes near itself (e.g. position-effect variegation). It is usually repetitive and forms structural functions such as centromeres or telomeres, in addition to acting as an attractor for other gene-expression or repression signals. Facultative hete ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Histone Deacetylase
Histone deacetylases (, HDAC) are a class of enzymes that remove acetyl groups (O=C-CH3) from an ε-N-acetyl lysine amino acid on a histone, allowing the histones to wrap the DNA more tightly. This is important because DNA is wrapped around histones, and DNA expression is regulated by acetylation and de-acetylation. Its action is opposite to that of histone acetyltransferase. HDAC proteins are now also called lysine deacetylases (KDAC), to describe their function rather than their target, which also includes non-histone proteins. HDAC super family Together with the acetylpolyamine amidohydrolases and the acetoin utilization proteins, the histone deacetylases form an ancient protein superfamily known as the histone deacetylase superfamily. Classes of HDACs in higher eukaryotes HDACs, are classified in four classes depending on sequence homology to the yeast original enzymes and domain organization: HDAC (except class III) contain zinc and are known as Zn2+-dependent hi ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Molecular Biology
Molecular biology is the branch of biology that seeks to understand the molecular basis of biological activity in and between cells, including biomolecular synthesis, modification, mechanisms, and interactions. The study of chemical and physical structure of biological macromolecules is known as molecular biology. Molecular biology was first described as an approach focused on the underpinnings of biological phenomena - uncovering the structures of biological molecules as well as their interactions, and how these interactions explain observations of classical biology. In 1945 the term molecular biology was used by physicist William Astbury. In 1953 Francis Crick, James Watson, Rosalind Franklin, and colleagues, working at Medical Research Council unit, Cavendish laboratory, Cambridge (now the MRC Laboratory of Molecular Biology), made a double helix model of DNA which changed the entire research scenario. They proposed the DNA structure based on previous research done by Ro ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Human Genome
The human genome is a complete set of nucleic acid sequences for humans, encoded as DNA within the 23 chromosome pairs in cell nuclei and in a small DNA molecule found within individual mitochondria. These are usually treated separately as the nuclear genome and the mitochondrial genome. Human genomes include both protein-coding DNA sequences and various types of DNA that does not encode proteins. The latter is a diverse category that includes DNA coding for non-translated RNA, such as that for ribosomal RNA, transfer RNA, ribozymes, small nuclear RNAs, and several types of regulatory RNAs. It also includes promoters and their associated gene-regulatory elements, DNA playing structural and replicatory roles, such as scaffolding regions, telomeres, centromeres, and origins of replication, plus large numbers of transposable elements, inserted viral DNA, non-functional pseudogenes and simple, highly-repetitive sequences. Introns make up a large percentage of non-coding DNA. ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
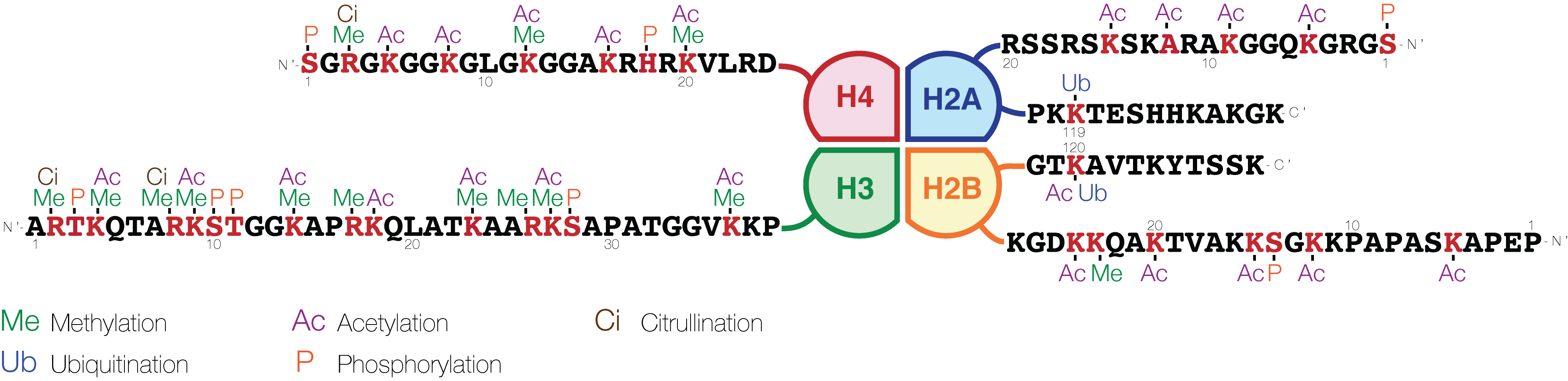
Histone Code
The histone code is a hypothesis that the transcription of genetic information encoded in DNA is in part regulated by chemical modifications (known as ''histone marks'') to histone proteins, primarily on their unstructured ends. Together with similar modifications such as DNA methylation it is part of the epigenetic code. Histones associate with DNA to form nucleosomes, which themselves bundle to form chromatin fibers, which in turn make up the more familiar chromosome. Histones are globular proteins with a flexible N-terminus (taken to be the tail) that protrudes from the nucleosome. Many of the histone tail modifications correlate very well to chromatin structure and both histone modification state and chromatin structure correlate well to gene expression levels. The critical concept of the histone code hypothesis is that the histone modifications serve to recruit other proteins by specific recognition of the modified histone via protein domains specialized for such purposes ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Epigenomics
Epigenomics is the study of the complete set of epigenetic modifications on the genetic material of a cell, known as the epigenome. The field is analogous to genomics and proteomics, which are the study of the genome and proteome of a cell. Epigenetic modifications are reversible modifications on a cell's DNA or histones that affect gene expression without altering the DNA sequence. Epigenomic maintenance is a continuous process and plays an important role in stability of eukaryotic genomes by taking part in crucial biological mechanisms like DNA repair. Plant flavones are said to be inhibiting epigenomic marks that cause cancers. Two of the most characterized epigenetic modifications are DNA methylation and histone modification. Epigenetic modifications play an important role in gene expression and regulation, and are involved in numerous cellular processes such as in differentiation/development and tumorigenesis. The study of epigenetics on a global level has been made possible o ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNA Methylation
DNA methylation is a biological process by which methyl groups are added to the DNA molecule. Methylation can change the activity of a DNA segment without changing the sequence. When located in a gene promoter, DNA methylation typically acts to repress gene transcription. In mammals, DNA methylation is essential for normal development and is associated with a number of key processes including genomic imprinting, X-chromosome inactivation, repression of transposable elements, aging, and carcinogenesis. As of 2016, two nucleobases have been found on which natural, enzymatic DNA methylation takes place: adenine and cytosine. The modified bases are N6-methyladenineD. B. Dunn, J. D. Smith: ''The occurrence of 6-methylaminopurine in deoxyribonucleic acids.'' In: ''Biochem J.'' 68(4), Apr 1958, S. 627–636. PMID 13522672. ., 5-methylcytosineB. F. Vanyushin, S. G. Tkacheva, A. N. Belozersky: ''Rare bases in animal DNA.'' In: ''Nature.'' 225, 1970, S. 948–949. PMID 4391887. and N4- ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Computational Epigenetics
Computational epigenetics uses statistical methods and mathematical modelling in epigenetic research. Due to the recent explosion of epigenome datasets, computational methods play an increasing role in all areas of epigenetic research. Definition Research in computational epigenetics comprises the development and application of bioinformatics methods for solving epigenetic questions, as well as computational data analysis and theoretical modeling in the context of epigenetics. This includes modelling of the effects of histone and DNA CpG island methylation. Current research areas Epigenetic data processing and analysis Various experimental techniques have been developed for genome-wide mapping of epigenetic information, the most widely used being ChIP-on-chip, ChIP-seq and bisulfite sequencing. All of these methods generate large amounts of data and require efficient ways of data processing and quality control by bioinformatic methods. Epigenome prediction A substantial amount ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Centromere
The centromere links a pair of sister chromatids together during cell division. This constricted region of chromosome connects the sister chromatids, creating a short arm (p) and a long arm (q) on the chromatids. During mitosis, spindle fibers attach to the centromere via the kinetochore. The physical role of the centromere is to act as the site of assembly of the kinetochores – a highly complex multiprotein structure that is responsible for the actual events of chromosome segregation – i.e. binding microtubules and signaling to the cell cycle machinery when all chromosomes have adopted correct attachments to the spindle, so that it is safe for cell division to proceed to completion and for cells to enter anaphase. There are, broadly speaking, two types of centromeres. "Point centromeres" bind to specific proteins that recognize particular DNA sequences with high efficiency. Any piece of DNA with the point centromere DNA sequence on it will typically form a centromere if pr ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Saccharomyces Cerevisiae
''Saccharomyces cerevisiae'' () (brewer's yeast or baker's yeast) is a species of yeast (single-celled fungus microorganisms). The species has been instrumental in winemaking, baking, and brewing since ancient times. It is believed to have been originally isolated from the skin of grapes. It is one of the most intensively studied eukaryotic model organisms in molecular biology, molecular and cell biology, much like ''Escherichia coli'' as the model bacteria, bacterium. It is the microorganism behind the most common type of fermentation (biochemistry), fermentation. ''S. cerevisiae'' cells are round to ovoid, 5–10 micrometre, μm in diameter. It reproduces by budding. Many proteins important in human biology were first discovered by studying their Homology (biology), homologs in yeast; these proteins include cell cycle proteins, signaling proteins, and protein-processing enzymes. ''S. cerevisiae'' is currently the only yeast cell known to have Berkeley body, Berkeley bo ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Yeast Mating
The yeast ''Saccharomyces cerevisiae'' is a simple microorganism, single-celled eukaryote with both a diploid and haploid mode of existence. The mating of yeast only occurs between haploids, which can be either the a or α (alpha) mating type and thus display simple sexual differentiation. Mating type is determined by a single locus (genetics), locus, ''MAT'', which in turn governs the sexual behaviour of both haploid and diploid cells. Through a form of genetic recombination, haploid yeast can switch mating type as often as every cell cycle. Mating type and the life cycle of ''Saccharomyces cerevisiae'' ''S. cerevisiae'' (yeast) can stably exist as either a diploid or a haploid. Both haploid and diploid yeast cells reproduce by mitosis, with daughter cells budding off of mother cells. Haploid cells are capable of mating with other haploid cells of the opposite mating type (an a cell can only mate with an α cell, and vice versa) to produce a stable diploid cell. Diploid cells, usu ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Sirtuin
Sirtuins are a family of signaling proteins involved in metabolic regulation. They are ancient in animal evolution and appear to possess a highly conserved structure throughout all kingdoms of life. Chemically, sirtuins are a class of proteins that possess either mono- ADP-ribosyltransferase or deacylase activity, including deacetylase, desuccinylase, demalonylase, demyristoylase and depalmitoylase activity. The name Sir2 comes from the yeast gene 'silent mating-type information regulation 2', the gene responsible for cellular regulation in yeast. From ''in vitro'' studies, sirtuins are implicated in influencing cellular processes like aging, transcription, apoptosis, inflammation and stress resistance, as well as energy efficiency and alertness during low-calorie situations. As of 2018, there was no clinical evidence that sirtuins affect human aging. Yeast Sir2 and some, but not all, sirtuins are protein deacetylases. Unlike other known protein deacetylases, which simply hyd ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |