|
PMP22
Growth arrest-specific protein 3 (GAS-3), also called peripheral myelin protein 22 (PMP22), is a protein which in humans is encoded by the ''PMP22'' gene. PMP22 is a 22 kDa transmembrane glycoprotein made up of 160 amino acids, and is mainly expressed in the Schwann cells of the peripheral nervous system. Schwann cells show high expression of PMP22, where it can constitute 2-5% of total protein content in compact myelin. Compact myelin is the bulk of the peripheral neuron's myelin sheath, a protective fatty layer that provides electrical insulation for the neuronal axon. The level of PMP22 expression is relatively low in the central nervous system of adults. Like other membrane proteins, newly translated PMP22 protein is temporarily sequestered to the endoplasmic reticulum (ER) and Golgi apparatus for post-translational modifications. PMP22 protein is glycosylated with an N terminus-linked sugar and co-localized with the chaperone protein calnexin in the ER. After the protein ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Myelin Protein Zero
Myelin protein zero (P0, MPZ) is a single membrane glycoprotein which in humans is encoded by the ''MPZ'' gene. P0 is a major structural component of the myelin sheath in the peripheral nervous system (PNS). Myelin protein zero is expressed by Schwann cells and accounts for over 50% of all proteins in the peripheral nervous system, making it the most common protein expressed in the PNS. Mutations in myelin protein zero can cause myelin deficiency and are associated with neuropathies like Charcot–Marie–Tooth disease and Dejerine–Sottas disease. Structure In humans, the gene that encodes myelin protein zero is located on chromosome 1 near the Duffy Locus or the Duffy Antigen/Chemokine Receptor. The gene is about 7,000 bases long and is divided into 6 exons. In total, myelin protein zero is 219 amino acids long and has many basic amino acid residues. Myelin protein zero consists of an extracellular N-terminal domain (amino acids 1–124), a single transmembrane region ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Chromosome 17
Chromosome 17 is one of the 23 pairs of chromosomes in humans. People normally have two copies of this chromosome. Chromosome 17 spans more than 83 million base pairs (the building material of DNA) and represents between 2.5 and 3% of the total DNA in cells. Chromosome 17 contains the Homeobox B gene cluster. Genes Number of genes The following are some of the gene count estimates of human chromosome 17. Because researchers use different approaches to genome annotation their predictions of the number of genes on each chromosome varies (for technical details, see gene prediction). Among various projects, the collaborative consensus coding sequence project ( CCDS) takes an extremely conservative strategy. So CCDS's gene number prediction represents a lower bound on the total number of human protein-coding genes. Gene list The following is a partial list of genes on human chromosome 17. For complete list, see the link in the infobox on the right. The following are some o ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, responding to stimuli, providing structure to cells and organisms, and transporting molecules from one location to another. Proteins differ from one another primarily in their sequence of amino acids, which is dictated by the nucleotide sequence of their genes, and which usually results in protein folding into a specific 3D structure that determines its activity. A linear chain of amino acid residues is called a polypeptide. A protein contains at least one long polypeptide. Short polypeptides, containing less than 20–30 residues, are rarely considered to be proteins and are commonly called peptides. The individual amino acid residues are bonded together by peptide bonds and adjacent amino acid residues. The sequence of amino acid residue ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Chaperone Protein
In molecular biology, molecular chaperones are proteins that assist the conformational folding or unfolding of large proteins or macromolecular protein complexes. There are a number of classes of molecular chaperones, all of which function to assist large proteins in proper protein folding during or after synthesis, and after partial denaturation. Chaperones are also involved in the translocation of proteins for proteolysis. The first molecular chaperones discovered were a type of assembly chaperones which assist in the assembly of nucleosomes from folded histones and DNA. One major function of molecular chaperones is to prevent the aggregation of misfolded proteins, thus many chaperone proteins are classified as heat shock proteins, as the tendency for protein aggregation is increased by heat stress. The majority of molecular chaperones do not convey any steric information for protein folding, and instead assist in protein folding by binding to and stabilizing folding intermediat ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Occludin
Occludin is an enzyme ( EC 1.6) that oxidizes NADH. It was first identified in epithelial cells as a 65 kDa integral plasma-membrane protein localized at the tight junctions. Together with Claudins, and zonula occludens-1 (ZO-1), occludin has been considered a staple of tight junctions, and although it was shown to regulate the formation, maintenance, and function of tight junctions, its precise mechanism of action remained elusive and most of its actions were initially attributed to conformational changes following selective phosphorylation, and its redox-sensitive dimerization. However, mounting evidence demonstrated that occludin is not only present in epithelial/endothelial cells, but is also expressed in large quantities in cells that do not have tight junctions but have very active metabolism: pericytes, neurons and astrocytes, oligodendrocytes, dendritic cells, monocytes/macrophages lymphocytes, and myocardium. Recent work, using molecular modeling, supported by biochemical ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Tight Junction Protein 1
Zonula occludens-1 ZO-1, also known as Tight junction protein-1 is a 220-kD peripheral membrane protein that is encoded by the ''TJP1'' gene in humans. It belongs to the family of ''zonula occludens proteins'' (ZO-1, ZO-2, and ZO-3), which are tight junction-associated proteins and of which, ZO-1 is the first to be cloned. It was first isolated in 1986 by Stevenson and Goodenough using a monoclonal antibody raised in rodent liver to recognise a 225-kD polypeptide in whole liver homogenates and in tight junction-enriched membrane fractions. It has a role as a scaffold protein which cross-links and anchors Tight Junction (TJ) strand proteins, which are fibril-like structures within the lipid bilayer, to the actin cytoskeleton. Function This gene encodes a protein located on a cytoplasmic membrane surface of intercellular tight junctions. The encoded protein may be involved in signal transduction at cell–cell junctions. Two transcript variants encoding distinct isoforms have bee ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Downregulation And Upregulation
In the biological context of organisms' production of gene products, downregulation is the process by which a cell decreases the quantity of a cellular component, such as RNA or protein, in response to an external stimulus. The complementary process that involves increases of such components is called upregulation. An example of downregulation is the cellular decrease in the expression of a specific receptor in response to its increased activation by a molecule, such as a hormone or neurotransmitter, which reduces the cell's sensitivity to the molecule. This is an example of a locally acting (negative feedback) mechanism. An example of upregulation is the response of liver cells exposed to such xenobiotic molecules as dioxin. In this situation, the cells increase their production of cytochrome P450 enzymes, which in turn increases degradation of these dioxin molecules. Downregulation or upregulation of an RNA or protein may also arise by an epigenetic alteration. Such an epigene ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Transmembrane Domains
A transmembrane domain (TMD) is a membrane-spanning protein domain. TMDs generally adopt an alpha helix topological conformation, although some TMDs such as those in porins can adopt a different conformation. Because the interior of the lipid bilayer is hydrophobic, the amino acid residues in TMDs are often hydrophobic, although proteins such as membrane pumps and ion channels can contain polar residues. TMDs vary greatly in length, sequence, and hydrophobicity, adopting organelle-specific properties. Functions of transmembrane domains Transmembrane domains are known to perform a variety of functions. These include: * Anchoring transmembrane proteins to the membrane. *Facilitating molecular transport of molecules such as ions and proteins across biological membranes; usually hydrophilic residues and binding sites in the TMDs help in this process. * Signal transduction across the membrane; many transmembrane proteins, such as G protein-coupled receptors, receive extracellula ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Alternative Splicing
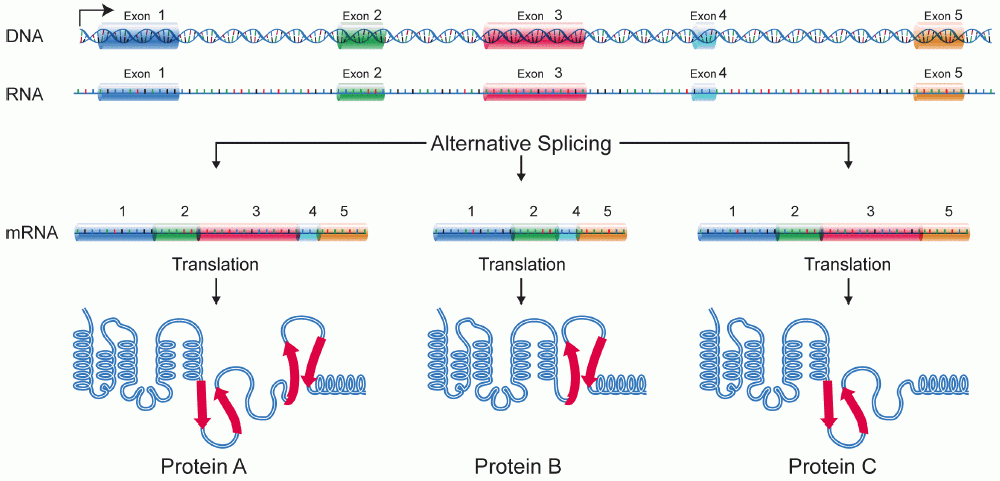
Alternative splicing, or alternative RNA splicing, or differential splicing, is an alternative splicing process during gene expression that allows a single gene to code for multiple proteins. In this process, particular exons of a gene may be included within or excluded from the final, processed messenger RNA (mRNA) produced from that gene. This means the exons are joined in different combinations, leading to different (alternative) mRNA strands. Consequently, the proteins translated from alternatively spliced mRNAs will contain differences in their amino acid sequence and, often, in their biological functions (see Figure). Biologically relevant alternative splicing occurs as a normal phenomenon in eukaryotes, where it increases the number of proteins that can be encoded by the genome. In humans, it is widely believed that ~95% of multi-exonic genes are alternatively spliced to produce functional alternative products from the same gene but many scientists believe that most o ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Post-transcriptional Modification
Transcriptional modification or co-transcriptional modification is a set of biological processes common to most eukaryotic cells by which an RNA primary transcript is chemically altered following transcription from a gene to produce a mature, functional RNA molecule that can then leave the nucleus and perform any of a variety of different functions in the cell. There are many types of post-transcriptional modifications achieved through a diverse class of molecular mechanisms. One example is the conversion of precursor messenger RNA transcripts into mature messenger RNA that is subsequently capable of being translated into protein. This process includes three major steps that significantly modify the chemical structure of the RNA molecule: the addition of a 5' cap, the addition of a 3' polyadenylated tail, and RNA splicing. Such processing is vital for the correct translation of eukaryotic genomes because the initial precursor mRNA produced by transcription often contains both exo ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Promoter (genetics)
In genetics, a promoter is a sequence of DNA to which proteins bind to initiate transcription of a single RNA transcript from the DNA downstream of the promoter. The RNA transcript may encode a protein (mRNA), or can have a function in and of itself, such as tRNA or rRNA. Promoters are located near the transcription start sites of genes, upstream on the DNA (towards the 5' region of the sense strand). Promoters can be about 100–1000 base pairs long, the sequence of which is highly dependent on the gene and product of transcription, type or class of RNA polymerase recruited to the site, and species of organism. Promoters control gene expression in bacteria and eukaryotes. RNA polymerase must attach to DNA near a gene for transcription to occur. Promoter DNA sequences provide an enzyme binding site. The -10 sequence is TATAAT. -35 sequences are conserved on average, but not in most promoters. Artificial promoters with conserved -10 and -35 elements transcribe more slowly. All D ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Coding Region
The coding region of a gene, also known as the coding sequence (CDS), is the portion of a gene's DNA or RNA that codes for protein. Studying the length, composition, regulation, splicing, structures, and functions of coding regions compared to non-coding regions over different species and time periods can provide a significant amount of important information regarding gene organization and evolution of prokaryotes and eukaryotes. This can further assist in mapping the human genome and developing gene therapy. Definition Although this term is also sometimes used interchangeably with exon, it is not the exact same thing: the exon is composed of the coding region as well as the 3' and 5' untranslated regions of the RNA, and so therefore, an exon would be partially made up of coding regions. The 3' and 5' untranslated regions of the RNA, which do not code for protein, are termed non-coding regions and are not discussed on this page. There is often confusion between coding region ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |



.png)