|
Messenger RNA Decapping
The process of messenger RNA decapping consists of hydrolysis of the 5' cap structure on the RNA exposing a 5' monophosphate. In eukaryotes, this 5' monophosphate is a substrate for the 5' exonuclease Xrn1 and the mRNA is quickly destroyed. There are many situations which may lead to the removal of the cap, some of which are discussed below. In prokaryotes, the initial mRNA transcript naturally possesses a 5'-triphosphate group after bacterial transcription; the enzyme RppH removes a pyrophosphate molecule from the 5' end, converting the 5'-triphosphate to a 5'-monophosphate, triggering mRNA degradation by ribonucleases. Translation and decay Inside cells, there is a balance between the processes of translation and mRNA decay. Messages which are being actively translated are bound by polysomes and the eukaryotic initiation factors eIF-4E and eIF-4G (in eukaryotes). This blocks access to the cap by the decapping enzyme DCP2 and protects the mRNA molecule. In nutrient-star ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Polysome
A polyribosome (or polysome or ergosome) is a group of ribosomes bound to an mRNA molecule like “beads” on a “thread”. It consists of a complex of an mRNA molecule and two or more ribosomes that act to translate mRNA instructions into polypeptides. Originally coined "ergosomes" in 1963, they were further characterized by Jonathan Warner, Paul M. Knopf, and Alex Rich Alexander Rich (15 November 1924 – 27 April 2015) was an American biologist and biophysicist. He was the William Thompson Sedgwick Professor of Biophysics at MIT (since 1958) and Harvard Medical School. Rich earned an A.B. ('' magna cum lau .... Polysomes are formed during the elongation phase when ribosomes and elongation factors synthesize the encoded polypeptide. Multiple ribosomes move along the coding region of mRNA, creating a polysome. The ability of multiple ribosomes to function on an mRNA molecule explains the limited abundance of mRNA in the cell. Polyribosome structure differs between pro ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Exosome Complex
The exosome complex (or PM/Scl complex, often just called the exosome) is a multi-protein intracellular complex capable of degrading various types of RNA (ribonucleic acid) molecules. Exosome complexes are found in both eukaryotic cells and archaea, while in bacteria a simpler complex called the degradosome carries out similar functions. The core of the exosome contains a six-membered ring structure to which other proteins are attached. In eukaryotic cells, the exosome complex is present in the cytoplasm, nucleus, and especially the nucleolus, although different proteins interact with the exosome complex in these compartments regulating the RNA degradation activity of the complex to substrates specific to these cell compartments. Substrates of the exosome include messenger RNA, ribosomal RNA, and many species of small RNAs. The exosome has an exoribonucleolytic function, meaning it degrades RNA starting at one end (the 3′ end in this case), and in eukaryotes also an endori ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Nonsense Mediated Decay
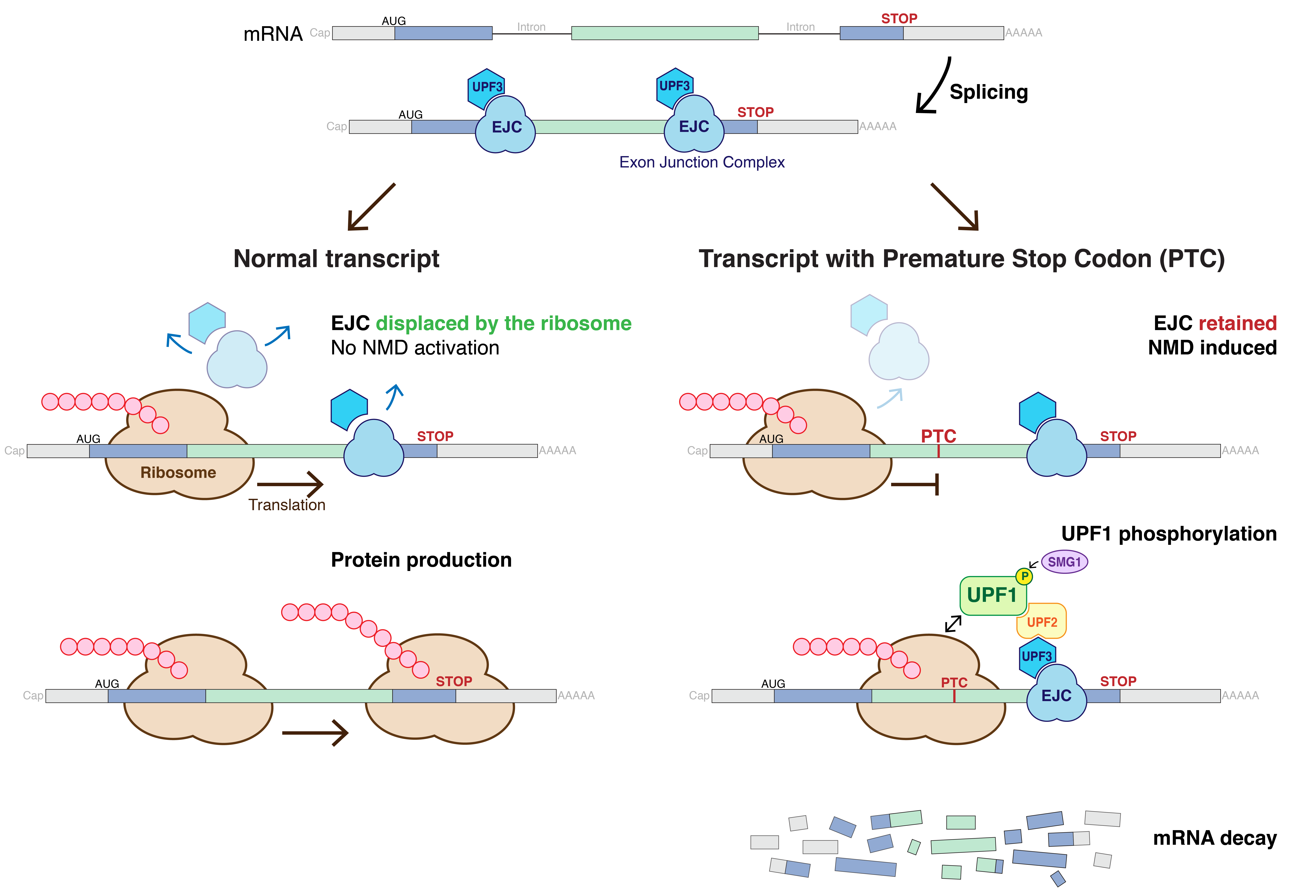
Nonsense-mediated mRNA decay (NMD) is a surveillance pathway that exists in all eukaryotes. Its main function is to reduce errors in gene expression by eliminating mRNA transcripts that contain premature stop codons. Translation of these aberrant mRNAs could, in some cases, lead to deleterious gain-of-function or dominant-negative activity of the resulting proteins. NMD was first described in human cells and in yeast almost simultaneously in 1979. This suggested broad phylogenetic conservation and an important biological role of this intriguing mechanism. NMD was discovered when it was realized that cells often contain unexpectedly low concentrations of mRNAs that are transcribed from alleles carrying nonsense mutations. Nonsense mutations code for a premature stop codon which causes the protein to be shortened. The truncated protein may or may not be functional, depending on the severity of what is not translated. In human genetics, NMD has the possibility to not only limit the ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
P-bodies
P-bodies, or processing bodies are distinct foci formed by phase separation within the cytoplasm of the eukaryotic cell consisting of many enzymes involved in mRNA turnover. P-bodies are highly conserved structures and have been observed in somatic cells originating from vertebrates and invertebrates, plants and yeast. To date, P-bodies have been demonstrated to play fundamental roles in general mRNA decay, nonsense-mediated mRNA decay, adenylate-uridylate-rich element mediated mRNA decay, and microRNA (miRNA) induced mRNA silencing. Not all mRNAs which enter P-bodies are degraded, as it has been demonstrated that some mRNAs can exit P-bodies and re-initiate translation. Purification and sequencing of the mRNA from purified processing bodies showed that these mRNAs are largely translationally repressed upstream of translation initiation and are protected from 5' mRNA decay. P-bodies are involved in decapping and degradation of unwanted mRNAs, storing mRNA until needed for tra ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DCP2
mRNA-decapping enzyme 2 is a protein that in humans is encoded by the ''DCP2'' gene In biology, the word gene (from , ; "... Wilhelm Johannsen coined the word gene to describe the Mendelian units of heredity..." meaning ''generation'' or ''birth'' or ''gender'') can have several different meanings. The Mendelian gene is a b .... DCP2 is a key component of an mRNA-decapping complex required for removal of the 5-prime cap from mRNA prior to its degradation from the 5-prime end (Fenger-Gron et al., 2005). upplied by OMIMref name="entrez" /> Interactions DCP2 has been shown to interact with DCP1A and UPF1. References Further reading * * * * * * * * * * * * * * {{gene-5-stub Nudix hydrolases ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
EIF-4G
Eukaryotic initiation factors (eIFs) are proteins or protein complexes involved in the initiation phase of eukaryotic translation. These proteins help stabilize the formation of ribosomal preinitiation complexes around the start codon and are an important input for post-transcription gene regulation. Several initiation factors form a complex with the small 40S ribosomal subunit and Met-tRNAiMet called the 43S preinitiation complex (43S PIC). Additional factors of the eIF4F complex (eIF4A, E, and G) recruit the 43S PIC to the five-prime cap structure of the mRNA, from which the 43S particle scans 5'-->3' along the mRNA to reach an AUG start codon. Recognition of the start codon by the Met-tRNAiMet promotes gated phosphate and eIF1 release to form the 48S preinitiation complex (48S PIC), followed by large 60S ribosomal subunit recruitment to form the 80S ribosome. There exist many more eukaryotic initiation factors than prokaryotic initiation factors, reflecting the greater biol ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
EIF-4E
Eukaryotic initiation factors (eIFs) are proteins or protein complexes involved in the initiation phase of eukaryotic translation. These proteins help stabilize the formation of ribosomal preinitiation complexes around the start codon and are an important input for post-transcription gene regulation. Several initiation factors form a complex with the small 40S ribosomal subunit and Met-tRNAiMet called the 43S preinitiation complex (43S PIC). Additional factors of the eIF4F complex (eIF4A, E, and G) recruit the 43S PIC to the five-prime cap structure of the mRNA, from which the 43S particle scans 5'-->3' along the mRNA to reach an AUG start codon. Recognition of the start codon by the Met-tRNAiMet promotes gated phosphate and eIF1 release to form the 48S preinitiation complex (48S PIC), followed by large 60S ribosomal subunit recruitment to form the 80S ribosome. There exist many more eukaryotic initiation factors than prokaryotic initiation factors, reflecting the greater biol ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Translation (genetics)
In molecular biology and genetics, translation is the process in which ribosomes in the cytoplasm or endoplasmic reticulum synthesize proteins after the process of transcription of DNA to RNA in the cell's nucleus. The entire process is called gene expression. In translation, messenger RNA (mRNA) is decoded in a ribosome, outside the nucleus, to produce a specific amino acid chain, or polypeptide. The polypeptide later folds into an active protein and performs its functions in the cell. The ribosome facilitates decoding by inducing the binding of complementary tRNA anticodon sequences to mRNA codons. The tRNAs carry specific amino acids that are chained together into a polypeptide as the mRNA passes through and is "read" by the ribosome. Translation proceeds in three phases: # Initiation: The ribosome assembles around the target mRNA. The first tRNA is attached at the start codon. # Elongation: The last tRNA validated by the small ribosomal subunit (''accommodation ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Messenger RNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of synthesizing a protein. mRNA is created during the process of transcription, where an enzyme (RNA polymerase) converts the gene into primary transcript mRNA (also known as pre-mRNA). This pre-mRNA usually still contains introns, regions that will not go on to code for the final amino acid sequence. These are removed in the process of RNA splicing, leaving only exons, regions that will encode the protein. This exon sequence constitutes mature mRNA. Mature mRNA is then read by the ribosome, and, utilising amino acids carried by transfer RNA (tRNA), the ribosome creates the protein. This process is known as translation. All of these processes form part of the central dogma of molecular biology, which describes the flow of genetic information in a biological system. As in DNA, genetic inf ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Pyrophosphate
In chemistry, pyrophosphates are phosphorus oxyanions that contain two phosphorus atoms in a P–O–P linkage. A number of pyrophosphate salts exist, such as disodium pyrophosphate (Na2H2P2O7) and tetrasodium pyrophosphate (Na4P2O7), among others. Often pyrophosphates are called diphosphates. The parent pyrophosphates are derived from partial or complete neutralization of pyrophosphoric acid. The pyrophosphate bond is also sometimes referred to as a phosphoanhydride bond, a naming convention which emphasizes the loss of water that occurs when two phosphates form a new P–O–P bond, and which mirrors the nomenclature for anhydrides of carboxylic acids. Pyrophosphates are found in ATP and other nucleotide triphosphates, which are important in biochemistry. The term pyrophosphate is also the name of esters formed by the condensation of a phosphorylated biological compound with inorganic phosphate, as for dimethylallyl pyrophosphate. This bond is also referred to as a high-energy ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Bacterial Transcription
Bacterial transcription is the process in which a segment of bacterial DNA is copied into a newly synthesized strand of messenger RNA (mRNA) with use of the enzyme RNA polymerase. The process occurs in three main steps: initiation, elongation, and termination; and the end result is a strand of mRNA that is complementary to a single strand of DNA. Generally, the transcribed region accounts for more than one gene. In fact, many prokaryotic genes occur in operons, which are a series of genes that work together to code for the same protein or gene product and are controlled by a single promoter. Bacterial RNA polymerase is made up of four subunits and when a fifth subunit attaches, called the σ-factor, the polymerase can recognize specific binding sequences in the DNA, called promoters. The binding of the σ-factor to the promoter is the first step in initiation. Once the σ-factor releases from the polymerase, elongation proceeds. The polymerase continues down the double stranded DN ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
.jpg)