|
List Of Biological Databases
Biological databases are stores of biological information. The journal ''Nucleic Acids Research'' regularly publishes special issues on biological databases and has a list of such databases. The 2018 issue has a list of about 180 such databases and updates to previously described databasesOmics Discovery Indexcan be used to browse and search several biological databases. Meta databases Meta databases are databases of databases that collect data about data to generate new data. They are capable of merging information from different sources and making it available in a new and more convenient form, or with an emphasis on a particular disease or organism.[metadatabase is a database model for metadata management, global query of independent database, and distributed data processing. The word metadatabase is an addition to the dictionary]. originally ,metadata was only common term referring simply to ''data about data '' such a tags ,keywords, and markup headers. * ConsensusPathDB: a ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Biological Databases
Biological databases are libraries of biological sciences, collected from scientific experiments, published literature, high-throughput experiment technology, and computational analysis. They contain information from research areas including genomics, proteomics, metabolomics, microarray gene expression, and phylogenetics. Information contained in biological databases includes gene function, structure, localization (both cellular and chromosomal), clinical effects of mutations as well as similarities of biological sequences and structures. Biological databases can be classified by the kind of data they collect (see below). Broadly, there are molecular databases (for sequences, molecules, etc.), functional databases (for physiology, enzyme activities, phenotypes, ecology etc), taxonomic databases (for species and other taxonomic ranks), images and other media, or specimens (for museum collections etc.) Databases are important tools in assisting scientists to analyze and explain ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
European Bioinformatics Institute
The European Bioinformatics Institute (EMBL-EBI) is an Intergovernmental Organization (IGO) which, as part of the European Molecular Biology Laboratory (EMBL) family, focuses on research and services in bioinformatics. It is located on the Wellcome Genome Campus in Hinxton near Cambridge, and employs over 600 full-time equivalent (FTE) staff. Institute leaders such as Rolf Apweiler, Alex Bateman, Ewan Birney, and Guy Cochrane, an adviser on the National Genomics Data Center Scientific Advisory Board, serve as part of the international research network of the BIG Data Center at the Beijing Institute of Genomics. Additionally, the EMBL-EBI hosts training programs that teach scientists the fundamentals of the work with biological data and promote the plethora of bioinformatic tools available for their research, both EMBL-EBI and non-EMBL-EBI-based. Bioinformatic services One of the roles of the EMBL-EBI is to index and maintain biological data in a set of databases, including E ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Model Organism
A model organism (often shortened to model) is a non-human species that is extensively studied to understand particular biological phenomena, with the expectation that discoveries made in the model organism will provide insight into the workings of other organisms. Model organisms are widely used to research human disease when human experimentation would be unfeasible or unethical. This strategy is made possible by the common descent of all living organisms, and the conservation of metabolic and developmental pathways and genetic material over the course of evolution. Studying model organisms can be informative, but care must be taken when generalizing from one organism to another. In researching human disease, model organisms allow for better understanding the disease process without the added risk of harming an actual human. The species chosen will usually meet a determined taxonomic equivalency to humans, so as to react to disease or its treatment in a way that resembles ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Digital Curation
Digital curation is the selection, preservation, maintenance, collection and archiving of digital assets. Digital curation establishes, maintains and adds value to repositories of digital data for present and future use. This is often accomplished by archivists, librarians, scientists, historians, and scholars. Enterprises are starting to use digital curation to improve the quality of information and data within their operational and strategic processes. Successful digital curation will mitigate digital obsolescence, keeping the information accessible to users indefinitely. Digital curation includes digital asset management, data curation, digital preservation, and electronic records management. Word History Much like the word ''archive'' has layered meanings and uses, the word ''curation'' is both a noun and a verb used originally in the field of museology to represent a wide range of activities, most often associated with collection care, long-term preservation, and exhibition ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Genome
In the fields of molecular biology and genetics, a genome is all the genetic information of an organism. It consists of nucleotide sequences of DNA (or RNA in RNA viruses). The nuclear genome includes protein-coding genes and non-coding genes, other functional regions of the genome such as regulatory sequences (see non-coding DNA), and often a substantial fraction of 'junk' DNA with no evident function. Almost all eukaryotes have mitochondria and a small mitochondrial genome. Algae and plants also contain chloroplasts with a chloroplast genome. The study of the genome is called genomics. The genomes of many organisms have been sequenced and various regions have been annotated. The International Human Genome Project reported the sequence of the genome for ''Homo sapiens'' in 200The Human Genome Project although the initial "finished" sequence was missing 8% of the genome consisting mostly of repetitive sequences. With advancements in technology that could handle sequenci ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Nucleosome Positioning Region Database
Nucleosome Positioning Region Database (NPRD) is a database of nucleosome formation sites (NFSs). See also References External links * http://srs6.bionet.nsc.ru/srs6/. Biological databases Genetics databases {{Biodatabase-stub ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
1000 Genomes Project
The 1000 Genomes Project (abbreviated as 1KGP), launched in January 2008, was an international research effort to establish by far the most detailed catalogue of human genetic variation. Scientists planned to sequence the genomes of at least one thousand anonymous participants from a number of different ethnic groups within the following three years, using newly developed technologies which were faster and less expensive. In 2010, the project finished its pilot phase, which was described in detail in a publication in the journal ''Nature''. In 2012, the sequencing of 1092 genomes was announced in a ''Nature'' publication. In 2015, two papers in ''Nature'' reported results and the completion of the project and opportunities for future research. Many rare variations, restricted to closely related groups, were identified, and eight structural-variation classes were analyzed. The project unites multidisciplinary research teams from institutes around the world, including China, Ita ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RefSeq
The Reference Sequence (RefSeq) database is an open access, annotated and curated collection of publicly available nucleotide sequences ( DNA, RNA) and their protein products. RefSeq was first introduced in 2000. This database is built by National Center for Biotechnology Information (NCBI), and, unlike GenBank, provides only a single record for each natural biological molecule (i.e. DNA, RNA or protein) for major organisms ranging from viruses to bacteria to eukaryotes. For each model organism, ''RefSeq'' aims to provide separate and linked records for the genomic DNA, the gene transcripts, and the proteins arising from those transcripts. ''RefSeq'' is limited to major organisms for which sufficient data are available (121,461 distinct "named" organisms as of July 2022), while GenBank includes sequences for any organism submitted (approximately 504,000 formally described species). RefSeq categories RefSeq collection comprises different data types, with different origins, so it i ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Online Mendelian Inheritance In Man
Online Mendelian Inheritance in Man (OMIM) is a continuously updated catalog of human genes and genetic disorders and traits, with a particular focus on the gene-phenotype relationship. , approximately 9,000 of the over 25,000 entries in OMIM represented phenotypes; the rest represented genes, many of which were related to known phenotypes. Versions and history OMIM is the online continuation of Dr. Victor A. McKusick's ''Mendelian Inheritance in Man'' (MIM), which was published in 12 editions between 1966 and 1998.McKusick, V. A. ''Mendelian Inheritance in Man. Catalogs of Autosomal Dominant, Autosomal Recessive and X-Linked Phenotypes.'' Baltimore, MD: Johns Hopkins University Press, 1st ed, 1996; 2nd ed, 1969; 3rd ed, 1971; 4th ed, 1975; 5th ed, 1978; 6th ed, 1983; 7th ed, 1986; 8th ed, 1988; 9th ed, 1990; 10th ed, 1992. Nearly all of the 1,486 entries in the first edition of MIM discussed phenotypes. MIM/OMIM is produced and curated at the Johns Hopkins School of Medicine ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
International HapMap Project
The International HapMap Project was an organization that aimed to develop a haplotype map (HapMap) of the human genome, to describe the common patterns of human genetic variation. HapMap is used to find genetic variants affecting health, disease and responses to drugs and environmental factors. The information produced by the project is made freely available for research. The International HapMap Project is a collaboration among researchers at academic centers, non-profit biomedical research groups and private companies in Canada, China (including Hong Kong), Japan, Nigeria, the United Kingdom, and the United States. It officially started with a meeting on October 27 to 29, 2002, and was expected to take about three years. It comprises two phases; the complete data obtained in Phase I were published on 27 October 2005. The analysis of the Phase II dataset was published in October 2007. The Phase III dataset was released in spring 2009 and the publication presenting the final resul ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
23andMe
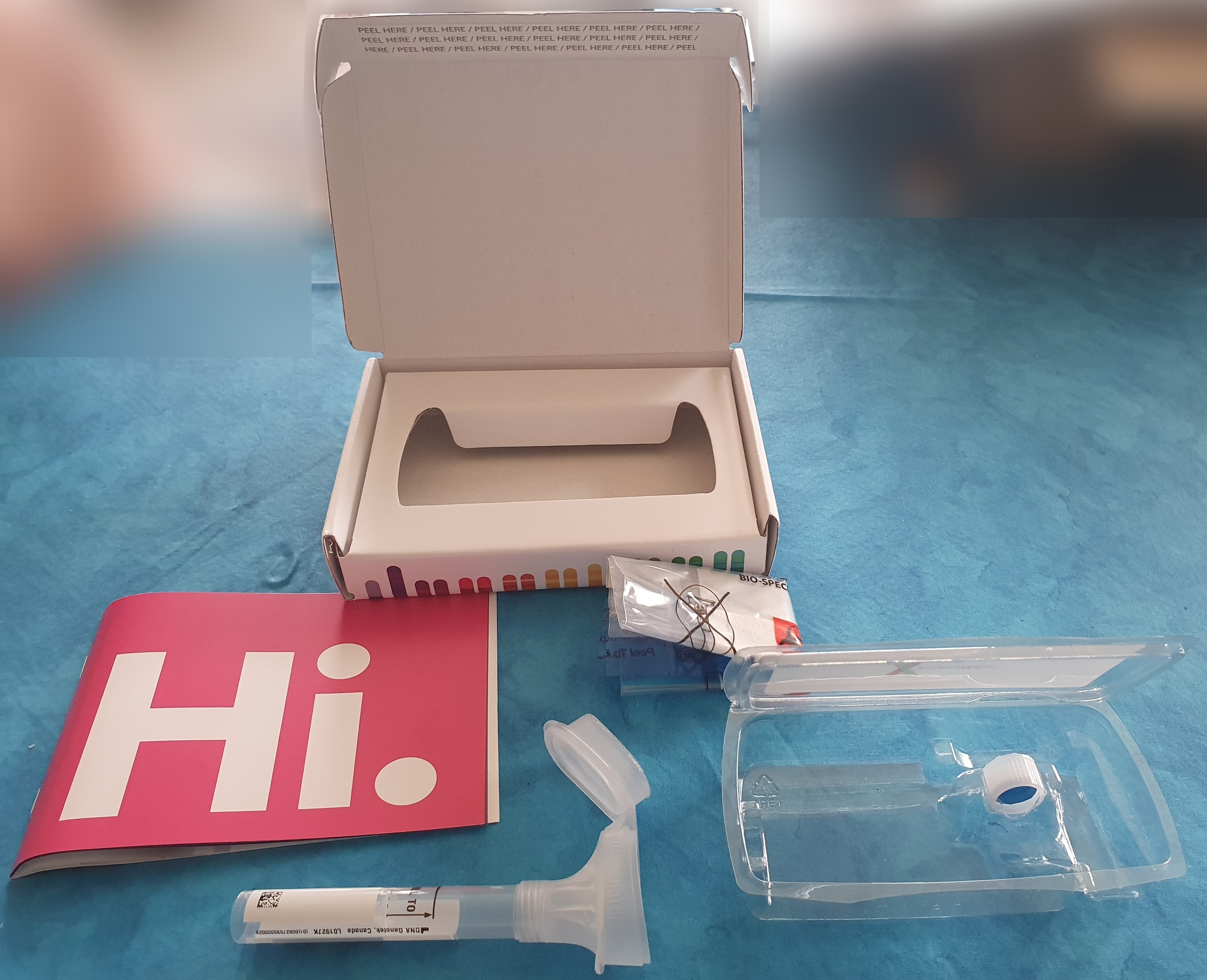
23andMe Holding Co. is a publicly held personal genomics and biotechnology company based in South San Francisco, California. It is best known for providing a direct-to-consumer genetic testing service in which customers provide a saliva sample that is laboratory analysed, using single nucleotide polymorphism genotyping, to generate reports relating to the customer's ancestry and genetic predispositions to health-related topics. The company's name is derived from the 23 pairs of chromosomes in a wild-type human cell. The company had a previously fraught relationship with the United States Food and Drug Administration (FDA) due to its genetic health tests; as of October 2015, DNA tests ordered in the US include a revised health component, per FDA approval. 23andMe has been selling a product with both ancestry and health-related components in Canada since October 2014, and in the UK since December 2014. In 2007, 23andMe became the first company to begin offering autosomal DNA tes ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Sequence Read Archive
The Sequence Read Archive (SRA, previously known as the Short Read Archive) is a bioinformatics database that provides a public repository for DNA sequencing data, especially the "short reads" generated by high-throughput sequencing, which are typically less than 1,000 base pairs in length. The archive is part of the International Nucleotide Sequence Database Collaboration (INSDC), and run as a collaboration between the NCBI, the European Bioinformatics Institute (EBI), and the DNA Data Bank of Japan (DDBJ). The archive was established by the National Center for Biotechnology Information (NCBI) in 2007 in order to provide a repository for data produced by RNA-Seq and ChIP-Seq studies as well as large-scale studies including the Human Microbiome Project and the 1000 Genomes Project. Originally called the Short Read Archive, the name was changed in anticipation of future sequencing technologies being able to produce longer sequence reads. The volume of data deposited in the Sequenc ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |