|
Ku86
Ku80 is a protein that, in humans, is encoded by the ''XRCC5'' gene. Together, Ku70 and Ku80 make up the Ku heterodimer, which binds to DNA double-strand break ends and is required for the non-homologous end joining (NHEJ) pathway of DNA repair. It is also required for V(D)J recombination, which utilizes the NHEJ pathway to promote antigen diversity in the mammalian immune system. In addition to its role in NHEJ, Ku is required for telomere length maintenance and subtelomeric gene silencing. Ku was originally identified when patients with systemic lupus erythematosus were found to have high levels of autoantibodies to the protein. Nomenclature Ku80 has been referred to by several names including: * Lupus Ku autoantigen protein p80 * ATP-dependent DNA helicase 2 subunit 2 * X-ray repair complementing defective repair in Chinese hamster cells 5 * X-ray repair cross-complementing 5 (XRCC5) Epigenetic repression The protein expression level of Ku80 can be repressed by epig ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ku (protein)
Ku is a dimeric protein complex that binds to DNA double-strand break ends and is required for the non-homologous end joining (NHEJ) pathway of DNA repair. Ku is evolutionarily conserved from bacteria to humans. The ancestral bacterial Ku is a homodimer (two copies of the same protein bound to each other). Eukaryotic Ku is a heterodimer of two polypeptides, Ku70 (XRCC6) and Ku80 (XRCC5), so named because the molecular weight of the human Ku proteins is around 70 kDa and 80 kDa. The two Ku subunits form a basket-shaped structure that threads onto the DNA end. Once bound, Ku can slide down the DNA strand, allowing more Ku molecules to thread onto the end. In higher eukaryotes, Ku forms a complex with the DNA-dependent protein kinase catalytic subunit (DNA-PKcs) to form the full DNA-dependent protein kinase, DNA-PK. Ku is thought to function as a molecular scaffold to which other proteins involved in NHEJ can bind, orienting the double-strand break for ligation. The Ku70 and ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNA Damage Theory Of Aging
The DNA damage theory of aging proposes that aging is a consequence of unrepaired accumulation of naturally occurring DNA damage. Damage in this context is a DNA alteration that has an abnormal structure. Although both mitochondrial and nuclear DNA damage can contribute to aging, nuclear DNA is the main subject of this analysis. Nuclear DNA damage can contribute to aging either indirectly (by increasing apoptosis or cellular senescence) or directly (by increasing cell dysfunction). Several review articles have shown that deficient DNA repair, allowing greater accumulation of DNA damage, causes premature aging; and that increased DNA repair facilitates greater longevity. Mouse models of nucleotide-excision–repair syndromes reveal a striking correlation between the degree to which specific DNA repair pathways are compromised and the severity of accelerated aging, strongly suggesting a causal relationship. Human population studies show that single-nucleotide polymorphisms in D ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Non-homologous End Joining
Non-homologous end joining (NHEJ) is a pathway that repairs double-strand breaks in DNA. NHEJ is referred to as "non-homologous" because the break ends are directly ligated without the need for a homologous template, in contrast to homology directed repair(HDR), which requires a homologous sequence to guide repair. NHEJ is active in both non-dividing and proliferating cells, while HDR is not readily accessible in non-dividing cells. The term "non-homologous end joining" was coined in 1996 by Moore and Haber. NHEJ is typically guided by short homologous DNA sequences called microhomologies. These microhomologies are often present in single-stranded overhangs on the ends of double-strand breaks. When the overhangs are perfectly compatible, NHEJ usually repairs the break accurately. Imprecise repair leading to loss of nucleotides can also occur, but is much more common when the overhangs are not compatible. Inappropriate NHEJ can lead to translocations and telomere fusion, hallmarks ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Carcinogenesis
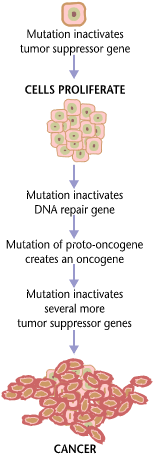
Carcinogenesis, also called oncogenesis or tumorigenesis, is the formation of a cancer, whereby normal cells are transformed into cancer cells. The process is characterized by changes at the cellular, genetic, and epigenetic levels and abnormal cell division. Cell division is a physiological process that occurs in almost all tissues and under a variety of circumstances. Normally, the balance between proliferation and programmed cell death, in the form of apoptosis, is maintained to ensure the integrity of tissues and organs. According to the prevailing accepted theory of carcinogenesis, the somatic mutation theory, mutations in DNA and epimutations that lead to cancer disrupt these orderly processes by interfering with the programming regulating the processes, upsetting the normal balance between proliferation and cell death. This results in uncontrolled cell division and the evolution of those cells by natural selection in the body. Only certain mutations lead to cancer w ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, responding to stimuli, providing structure to cells and organisms, and transporting molecules from one location to another. Proteins differ from one another primarily in their sequence of amino acids, which is dictated by the nucleotide sequence of their genes, and which usually results in protein folding into a specific 3D structure that determines its activity. A linear chain of amino acid residues is called a polypeptide. A protein contains at least one long polypeptide. Short polypeptides, containing less than 20–30 residues, are rarely considered to be proteins and are commonly called peptides. The individual amino acid residues are bonded together by peptide bonds and adjacent amino acid residues. The sequence of amino acid residue ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
POU2F1
POU domain, class 2, transcription factor 1 is a protein that in humans is encoded by the ''POU2F1'' gene. Interactions POU2F1 has been shown to interact with: * EPRS, * Glucocorticoid receptor, * Glyceraldehyde 3-phosphate dehydrogenase, * Host cell factor C1, * Ku80, * MNAT1 * NPAT, * Nuclear receptor co-repressor 2, * POU2AF1, * RELA, * Retinoid X receptor alpha, * SNAPC4, * Sp1 transcription factor, and * TATA binding protein. See also * Octamer transcription factor Octamer transcription factors are a protein family, family of transcription factors which binds to the "ATTTGCAT" Nucleic acid sequence, DNA sequence. Their DNA-binding domain is a POU domain. There are eight Octamer proteins in humans (Oct1–11) ... References Further reading * * * * * * * * * * * * * * * * * * External links * {{Transcription factors, g3 POU-domain proteins ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
PCNA
Proliferating cell nuclear antigen (PCNA) is a DNA clamp that acts as a processivity factor for DNA polymerase δ in eukaryotic cells and is essential for replication. PCNA is a homotrimer and achieves its processivity by encircling the DNA, where it acts as a scaffold to recruit proteins involved in DNA replication, DNA repair, chromatin remodeling and epigenetics. Many proteins interact with PCNA via the two known PCNA-interacting motifs PCNA-interacting peptide (PIP) box and AlkB homologue 2 PCNA interacting motif (APIM). Proteins binding to PCNA via the PIP-box are mainly involved in DNA replication whereas proteins binding to PCNA via APIM are mainly important in the context of genotoxic stress. Function The protein encoded by this gene is found in the nucleus and is a cofactor of DNA polymerase delta. The encoded protein acts as a homotrimer and helps increase the processivity of leading strand synthesis during DNA replication. In response to DNA damage, this protein is ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
NCOA6
Nuclear receptor coactivator 6 is a protein that in humans is encoded by the ''NCOA6'' gene. Function The protein encoded by this gene is a transcriptional coactivator that can interact with nuclear hormone receptors to enhance their transcriptional activator functions. The encoded protein has been shown to be involved in the hormone-dependent coactivation of several receptors, including prostanoid, retinoid, vitamin D3, thyroid hormone, and steroid receptors. The encoded protein may also act as a general coactivator since it has been shown to interact with some basal transcription factors, histone acetyltransferases, and methyltransferases. Interactions NCOA6 has been shown to interact with: * ASCL2 and * Activating transcription factor 2, * Androgen receptor, * CREB-binding protein, * DNA-PKcs, * E2F1, * EP300, * Estrogen receptor alpha, * Estrogen receptor beta, * HBXIP, * HIST2H3C, * HSF1, * Ku70, * Ku80, * Liver X receptor beta, * MLL3, * RBBP5, * Reti ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
GCN5L2
__NOTOC__ Histone acetyltransferase KAT2A is an enzyme that in humans is encoded by the ''KAT2A'' gene. Interactions GCN5L2 has been shown to interact with: * DDB1, * Ku70, * Ku80, * TADA2L, * TAF9, and * Transcription initiation protein SPT3 homolog Transcription initiation protein SPT3 homolog is a protein that in humans is encoded by the ''SUPT3H'' gene. Interactions Transcription initiation protein SPT3 homolog has been shown to interact with GCN5L2, TAF6L, TADA3L, TAF5L, SF3B3, SUPT7L .... References Further reading * * * * * * * * * * * * * * * * * * * External links * {{gene-17-stub ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNA-PKcs
DNA-dependent protein kinase, catalytic subunit, also known as DNA-PKcs, is an enzyme that in humans is encoded by the gene designated as ''PRKDC'' or ''XRCC7''. DNA-PKcs belongs to the phosphatidylinositol 3-kinase-related kinase protein family. The DNA-Pkcs protein is a serine/threonine protein kinase comprising a single polypeptide chain of 4,128 amino acids. Function DNA-PKcs is the catalytic subunit of a nuclear DNA-dependent serine/threonine protein kinase called DNA-PK. The second component is the autoimmune antigen Ku. On its own, DNA-PKcs is inactive and relies on Ku to direct it to DNA ends and trigger its kinase activity. DNA-PKcs is required for the non-homologous end joining (NHEJ) pathway of DNA repair, which rejoins double-strand breaks. It is also required for V(D)J recombination, a process that utilizes NHEJ to promote immune system diversity. DNA-PKcs knockout mice have severe combined immunodeficiency due to their V(D)J recombination defect. Many proteins ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cancer Epigenetics
Cancer epigenetics is the study of epigenetics, epigenetic modifications to the DNA of cancer cells that do not involve a change in the nucleotide sequence, but instead involve a change in the way the genetic code is expressed. Epigenetic mechanisms are necessary to maintain normal sequences of tissue specific gene expression and are crucial for normal development. They may be just as important, if not even more important, than mutation, genetic mutations in a cell's transformation to cancer. The disturbance of epigenetic processes in cancers, can lead to a loss of gene expression, expression of genes that occurs about 10 times more frequently by transcription silencing (caused by epigenetic promoter hypermethylation of CpG site#Methylation, silencing, cancer, and aging, CpG islands) than by mutations. As Vogelstein et al. points out, in a colorectal cancer there are usually about 3 to 6 driver mutations and 33 to 66 Genetic hitchhiking, hitchhiker or passenger mutations. However, in ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |