|
Intronerator
The Intronerator is a database of alternatively spliced genes and a database of introns for Caenorhabditis elegans. See also * Alternative splicing * AspicDB * EDAS * Hollywood (database) * List of biological databases Biological databases are stores of biological information. The journal ''Nucleic Acids Research'' regularly publishes special issues on biological databases and has a list of such databases. The 2018 issue has a list of about 180 such databases an ... References External links A working copy of the Intronerator no longer exists as of 2003. Equivalent functions can be performed with the U.C. Santa Cruz genome browser: * http://genome.ucsc.edu/cgi-bin/hgTracks?clade=worm&organism=C._elegans Biological databases Gene expression Spliceosome RNA splicing {{Biodatabase-stub ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Alternative Splicing
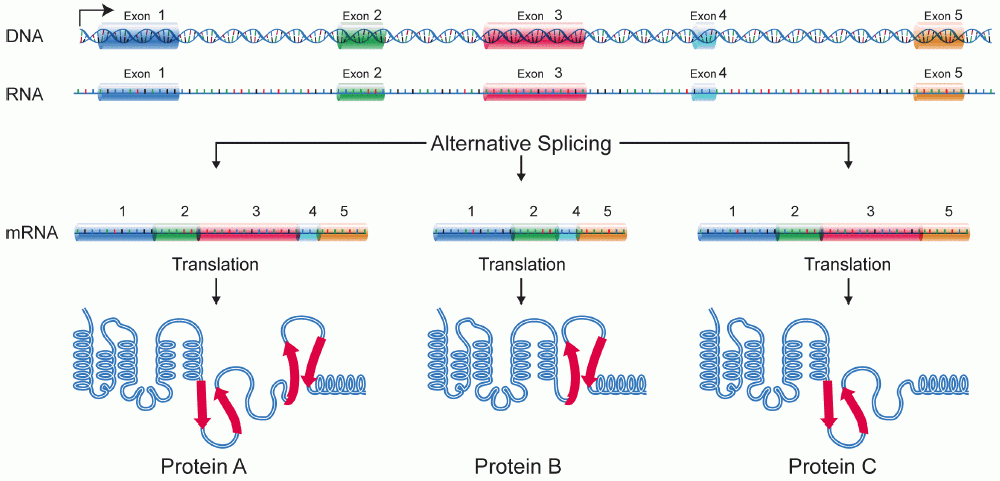
Alternative splicing, or alternative RNA splicing, or differential splicing, is an alternative splicing process during gene expression that allows a single gene to code for multiple proteins. In this process, particular exons of a gene may be included within or excluded from the final, processed messenger RNA (mRNA) produced from that gene. This means the exons are joined in different combinations, leading to different (alternative) mRNA strands. Consequently, the proteins translated from alternatively spliced mRNAs will contain differences in their amino acid sequence and, often, in their biological functions (see Figure). Biologically relevant alternative splicing occurs as a normal phenomenon in eukaryotes, where it increases the number of proteins that can be encoded by the genome. In humans, it is widely believed that ~95% of multi-exonic genes are alternatively spliced to produce functional alternative products from the same gene but many scientists believe that most o ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Caenorhabditis Elegans
''Caenorhabditis elegans'' () is a free-living transparent nematode about 1 mm in length that lives in temperate soil environments. It is the type species of its genus. The name is a blend of the Greek ''caeno-'' (recent), ''rhabditis'' (rod-like) and Latin ''elegans'' (elegant). In 1900, Maupas initially named it '' Rhabditides elegans.'' Osche placed it in the subgenus ''Caenorhabditis'' in 1952, and in 1955, Dougherty raised ''Caenorhabditis'' to the status of genus. ''C. elegans'' is an unsegmented pseudocoelomate and lacks respiratory or circulatory systems. Most of these nematodes are hermaphrodites and a few are males. Males have specialised tails for mating that include spicules. In 1963, Sydney Brenner proposed research into ''C. elegans,'' primarily in the area of neuronal development. In 1974, he began research into the molecular and developmental biology of ''C. elegans'', which has since been extensively used as a model organism. It was the first multicellu ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Alternative Splicing
Alternative splicing, or alternative RNA splicing, or differential splicing, is an alternative splicing process during gene expression that allows a single gene to code for multiple proteins. In this process, particular exons of a gene may be included within or excluded from the final, processed messenger RNA (mRNA) produced from that gene. This means the exons are joined in different combinations, leading to different (alternative) mRNA strands. Consequently, the proteins translated from alternatively spliced mRNAs will contain differences in their amino acid sequence and, often, in their biological functions (see Figure). Biologically relevant alternative splicing occurs as a normal phenomenon in eukaryotes, where it increases the number of proteins that can be encoded by the genome. In humans, it is widely believed that ~95% of multi-exonic genes are alternatively spliced to produce functional alternative products from the same gene but many scientists believe that most o ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Database
In computing, a database is an organized collection of data stored and accessed electronically. Small databases can be stored on a file system, while large databases are hosted on computer clusters or cloud storage. The design of databases spans formal techniques and practical considerations, including data modeling, efficient data representation and storage, query languages, security and privacy of sensitive data, and distributed computing issues, including supporting concurrent access and fault tolerance. A database management system (DBMS) is the software that interacts with end users, applications, and the database itself to capture and analyze the data. The DBMS software additionally encompasses the core facilities provided to administer the database. The sum total of the database, the DBMS and the associated applications can be referred to as a database system. Often the term "database" is also used loosely to refer to any of the DBMS, the database system or an appli ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Jim Kent
William James Kent (born February 10, 1960) is an American research scientist and computer programmer. He has been a contributor to genome database projects and the 2003 winner of the Benjamin Franklin Award. Early life Kent was born in Hawaii and grew up in San Francisco, California, United States. Computer animation Kent began his programming career in 1983 with Island Graphics Inc. where he wrote the Aegis Animator program for the Amiga home computer. This program combined polygon tweening in 3D with simple 2D cel-based animation. In 1985 he founded and ran a software company, Dancing Flame, which adapted the Aegis Animator to the Atari ST, and created Cyber Paint for that machine. Cyber Paint was a 2D animation program that brought together a wide variety of animation and paint functionality and the delta-compressed animation format developed for CAD-3D. The user could move freely between animation frames and paint arbitrarily, or utilize various animation tools for a ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
EDAS
EDAS was a database of alternatively spliced Alternative splicing, or alternative RNA splicing, or differential splicing, is an alternative splicing process during gene expression that allows a single gene to code for multiple proteins. In this process, particular exons of a gene may be ... human genes. It doesn't seem to exist anymore. See also * AspicDB database References External links * http://www.gene-bee.msu.ru/edas/. Genetics databases Gene expression Spliceosome RNA splicing {{Biodatabase-stub ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Hollywood (database)
Hollywood is a RNA splicing database containing data for the splicing of orthologous genes in different species. See also * Alternative splicing * EDAS EDAS was a database of alternatively spliced Alternative splicing, or alternative RNA splicing, or differential splicing, is an alternative splicing process during gene expression that allows a single gene to code for multiple proteins. In t ... * AspicDB References External links * http://hollywood.mit.edu Genetics databases Gene expression Spliceosome RNA splicing {{Biodatabase-stub ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
List Of Biological Databases
Biological databases are stores of biological information. The journal ''Nucleic Acids Research'' regularly publishes special issues on biological databases and has a list of such databases. The 2018 issue has a list of about 180 such databases and updates to previously described databasesOmics Discovery Indexcan be used to browse and search several biological databases. Meta databases Meta databases are databases of databases that collect data about data to generate new data. They are capable of merging information from different sources and making it available in a new and more convenient form, or with an emphasis on a particular disease or organism.[metadatabase is a database model for metadata management, global query of independent database, and distributed data processing. The word metadatabase is an addition to the dictionary]. originally ,metadata was only common term referring simply to ''data about data '' such a tags ,keywords, and markup headers. * ConsensusPathDB: a ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Biological Databases
Biological databases are libraries of biological sciences, collected from scientific experiments, published literature, high-throughput experiment technology, and computational analysis. They contain information from research areas including genomics, proteomics, metabolomics, microarray gene expression, and phylogenetics. Information contained in biological databases includes gene function, structure, localization (both cellular and chromosomal), clinical effects of mutations as well as similarities of biological sequences and structures. Biological databases can be classified by the kind of data they collect (see below). Broadly, there are molecular databases (for sequences, molecules, etc.), functional databases (for physiology, enzyme activities, phenotypes, ecology etc), taxonomic databases (for species and other taxonomic ranks), images and other media, or specimens (for museum collections etc.) Databases are important tools in assisting scientists to analyze and explain ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Gene Expression
Gene expression is the process by which information from a gene is used in the synthesis of a functional gene product that enables it to produce end products, protein or non-coding RNA, and ultimately affect a phenotype, as the final effect. These products are often proteins, but in non-protein-coding genes such as transfer RNA (tRNA) and small nuclear RNA (snRNA), the product is a functional non-coding RNA. Gene expression is summarized in the central dogma of molecular biology first formulated by Francis Crick in 1958, further developed in his 1970 article, and expanded by the subsequent discoveries of reverse transcription and RNA replication. The process of gene expression is used by all known life—eukaryotes (including multicellular organisms), prokaryotes (bacteria and archaea), and utilized by viruses—to generate the macromolecular machinery for life. In genetics, gene expression is the most fundamental level at which the genotype gives rise to the phenotype, '' ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Spliceosome
A spliceosome is a large ribonucleoprotein (RNP) complex found primarily within the nucleus of eukaryotic cells. The spliceosome is assembled from small nuclear RNAs (snRNA) and numerous proteins. Small nuclear RNA (snRNA) molecules bind to specific proteins to form a small nuclear ribonucleoprotein complex (snRNP, pronounced “snurps”), which in turn combines with other snRNPs to form a large ribonucleoprotein complex called a spliceosome. The spliceosome removes introns from a transcribed pre-mRNA, a type of primary transcript. This process is generally referred to as splicing. An analogy is a film editor, who selectively cuts out irrelevant or incorrect material (equivalent to the introns) from the initial film and sends the cleaned-up version to the director for the final cut. However, sometimes the RNA within the intron acts as a ribozyme, splicing itself without the use of a spliceosome or protein enzymes. History In 1977, work by the Sharp and Roberts labs reveale ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |