|
Genotype-first Approach
The genotype-first approach is a type of strategy used in genetic epidemiological studies to associate specific genotypes to apparent clinical phenotypes of a complex disease or trait. As opposed to “phenotype-first”, the traditional strategy that has been guiding genome-wide association studies (GWAS) so far, this approach characterizes individuals first by a statistically common genotype based on molecular tests prior to clinical phenotypic classification. This method of grouping leads to patient evaluations based on a shared genetic etiology for the observed phenotypes, regardless of their suspected diagnosis. Thus, this approach can prevent initial phenotypic bias and allow for identification of genes that pose a significant contribution to the disease etiology.Stessman, H. A., Bernier, R. & Eichler, E. E. A genotype-first approach to defining the subtypes of a complex disease. Cell 156, 872–877 (2014).Mefford, H. C. Genotype to phenotype-discovery and characteriz ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Genotype-first Approach
The genotype-first approach is a type of strategy used in genetic epidemiological studies to associate specific genotypes to apparent clinical phenotypes of a complex disease or trait. As opposed to “phenotype-first”, the traditional strategy that has been guiding genome-wide association studies (GWAS) so far, this approach characterizes individuals first by a statistically common genotype based on molecular tests prior to clinical phenotypic classification. This method of grouping leads to patient evaluations based on a shared genetic etiology for the observed phenotypes, regardless of their suspected diagnosis. Thus, this approach can prevent initial phenotypic bias and allow for identification of genes that pose a significant contribution to the disease etiology.Stessman, H. A., Bernier, R. & Eichler, E. E. A genotype-first approach to defining the subtypes of a complex disease. Cell 156, 872–877 (2014).Mefford, H. C. Genotype to phenotype-discovery and characteriz ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Mutation
In biology, a mutation is an alteration in the nucleic acid sequence of the genome of an organism, virus, or extrachromosomal DNA. Viral genomes contain either DNA or RNA. Mutations result from errors during DNA or viral replication, mitosis, or meiosis or other types of damage to DNA (such as pyrimidine dimers caused by exposure to ultraviolet radiation), which then may undergo error-prone repair (especially microhomology-mediated end joining), cause an error during other forms of repair, or cause an error during replication (translesion synthesis). Mutations may also result from insertion or deletion of segments of DNA due to mobile genetic elements. Mutations may or may not produce detectable changes in the observable characteristics (phenotype) of an organism. Mutations play a part in both normal and abnormal biological processes including: evolution, cancer, and the development of the immune system, including junctional diversity. Mutation is the ultimate source o ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Whole-genome Sequencing
Whole genome sequencing (WGS), also known as full genome sequencing, complete genome sequencing, or entire genome sequencing, is the process of determining the entirety, or nearly the entirety, of the DNA sequence of an organism's genome at a single time. This entails sequencing all of an organism's chromosomal DNA as well as DNA contained in the mitochondria and, for plants, in the chloroplast. Whole genome sequencing has largely been used as a research tool, but was being introduced to clinics in 2014. In the future of personalized medicine, whole genome sequence data may be an important tool to guide therapeutic intervention. The tool of gene sequencing at SNP level is also used to pinpoint functional variants from association studies and improve the knowledge available to researchers interested in evolutionary biology, and hence may lay the foundation for predicting disease susceptibility and drug response. Whole genome sequencing should not be confused with DNA profilin ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Microarray
A microarray is a multiplex lab-on-a-chip. Its purpose is to simultaneously detect the expression of thousands of genes from a sample (e.g. from a tissue). It is a two-dimensional array on a solid substrate—usually a glass slide or silicon thin-film cell—that assays (tests) large amounts of biological material using high-throughput screening miniaturized, multiplexed and parallel processing and detection methods. The concept and methodology of microarrays was first introduced and illustrated in antibody microarrays (also referred to as antibody matrix) by Tse Wen Chang in 1983 in a scientific publication and a series of patents. The "gene chip" industry started to grow significantly after the 1995 ''Science Magazine'' article by the Ron Davis and Pat Brown labs at Stanford University. With the establishment of companies, such as Affymetrix, Agilent, Applied Microarrays, Arrayjet, Illumina, and others, the technology of DNA microarrays has become the most sophisticated and ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
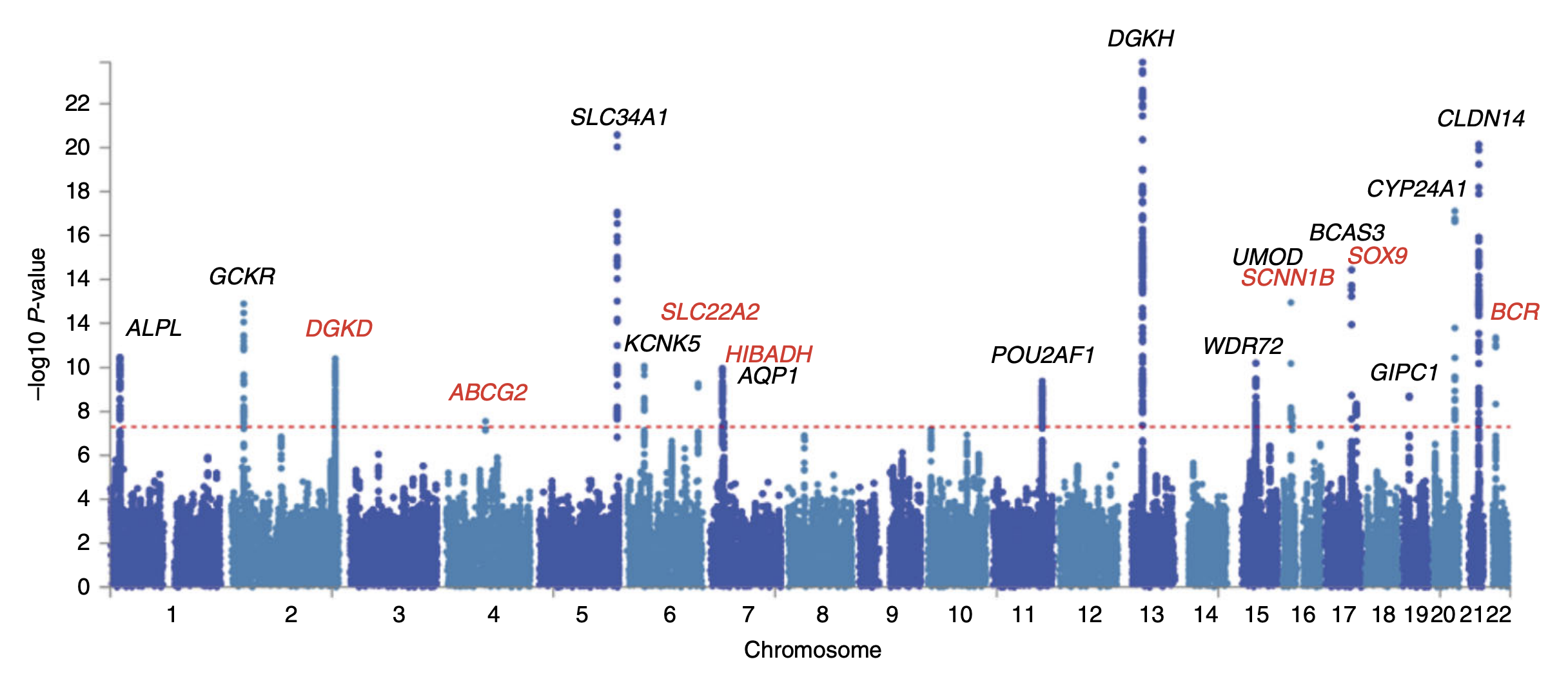
Genome-wide Association Study
In genomics, a genome-wide association study (GWA study, or GWAS), also known as whole genome association study (WGA study, or WGAS), is an observational study of a genome-wide set of Single-nucleotide polymorphism, genetic variants in different individuals to see if any variant is associated with a trait. GWA studies typically focus on associations between single-nucleotide polymorphisms (SNPs) and traits like major human diseases, but can equally be applied to any other genetic variants and any other organisms. When applied to human data, GWA studies compare the DNA of participants having varying phenotypes for a particular trait or disease. These participants may be people with a disease (cases) and similar people without the disease (controls), or they may be people with different phenotypes for a particular trait, for example blood pressure. This approach is known as phenotype-first, in which the participants are classified first by their clinical manifestation(s), as oppose ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Genetic Disorder
A genetic disorder is a health problem caused by one or more abnormalities in the genome. It can be caused by a mutation in a single gene (monogenic) or multiple genes (polygenic) or by a chromosomal abnormality. Although polygenic disorders are the most common, the term is mostly used when discussing disorders with a single genetic cause, either in a gene or chromosome. The mutation responsible can occur spontaneously before embryonic development (a ''de novo'' mutation), or it can be Heredity, inherited from two parents who are carriers of a faulty gene (autosomal recessive inheritance) or from a parent with the disorder (autosomal dominant inheritance). When the genetic disorder is inherited from one or both parents, it is also classified as a hereditary disease. Some disorders are caused by a mutation on the X chromosome and have X-linked inheritance. Very few disorders are inherited on the Y linkage, Y chromosome or Mitochondrial disease#Causes, mitochondrial DNA (due to t ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Variants Of Unknown Significance
A variant of uncertain (or unknown) significance (VUS) is a genetic variant that has been identified through genetic testing but whose significance to the function or health of an organism is not known. Two related terms are "gene of uncertain significance" (GUS), which refers to a gene that has been identified through genome sequencing but whose connection to a human disease has not been established, and "insignificant mutation", referring to a gene variant that has no impact on the health or function of an organism. The term "variant' is favored in clinical practice over "mutation" because it can be used to describe an allele more precisely (i.e. without inherently connoting pathogenicity). When the variant has no impact on health, it is called a "benign variant". When it is associated with a disease, it is called a "pathogenic variant". A "pharmacogenomic variant" has an effect only when an individual takes a particular drug and therefore is neither benign nor pathogenic. A VUS i ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Developmental Disability
Developmental disability is a diverse group of chronic conditions, comprising mental or physical impairments that arise before adulthood. Developmental disabilities cause individuals living with them many difficulties in certain areas of life, especially in "language, mobility, learning, self-help, and independent living".Center for Disease Control and Prevention. (2013)Developmental disabilities.Retrieved October 18, 2013 Developmental disabilities can be detected early on and persist throughout an individual's lifespan. Developmental disability that affects all areas of a child's development is sometimes referred to as global developmental delay. The most common developmental disabilities are: * Motor disorders, and learning difficulties such as dyslexia, Tourette's syndrome, dyspraxia, dysgraphia, Irlen syndrome, and dyscalculia. * Autism and Asperger syndrome are a series of conditions called autistic spectrum disorders that causes difficulties in communications. Autistic s ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Lim Et Al
Lim or LIM may refer to: Name * Lim (Korean surname), a common Korean surname * Lim (Chinese surname), Hokkien, Hakka, Teochew and Hainanese spelling of the Chinese family name "Lin" * Liza Lim (born 1966), Australian classical composer Abbreviations * Lanes in metres, a unit of measure for vehicle ferries * LIM College (Laboratory Institute of Merchandising), New York City, US * Linear induction motor * Logical Information Machines, Chicago, US software company * LIM domain, a protein-protein interaction domain * Lotus-Intel-Microsoft, the alliance responsible for the Expanded Memory Specification (EMS) Places * IATA airport code for Jorge Chávez International Airport, Lima, Peru) * Lim (Croatia), a bay and a valley * Lim (river), in Montenegro, Albania, Bosnia and Herzegovina and Serbia * Lim Island or Adır Island, Lake Van, Turkey * Lim, Bắc Ninh, a township in Vietnam Others * A symbol for the limit (mathematics) operator * Lim (musical instrument), a Bhutanese flute ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
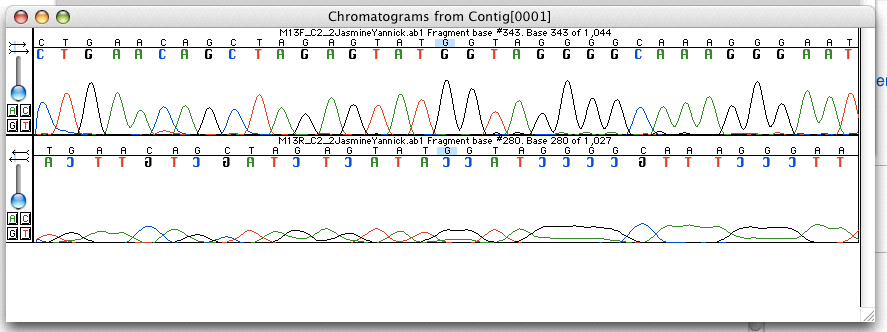
Sanger Sequencing
Sanger sequencing is a method of DNA sequencing that involves electrophoresis and is based on the random incorporation of chain-terminating dideoxynucleotides by DNA polymerase during in vitro DNA replication. After first being developed by Frederick Sanger and colleagues in 1977, it became the most widely used sequencing method for approximately 40 years. It was first commercialized by Applied Biosystems in 1986. More recently, higher volume Sanger sequencing has been replaced by next generation sequencing methods, especially for large-scale, automated genome analyses. However, the Sanger method remains in wide use for smaller-scale projects and for validation of deep sequencing results. It still has the advantage over short-read sequencing technologies (like Illumina) in that it can produce DNA sequence reads of > 500 nucleotides and maintains a very low error rate with accuracies around 99.99%. Sanger sequencing is still actively being used in efforts for public health initiative ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
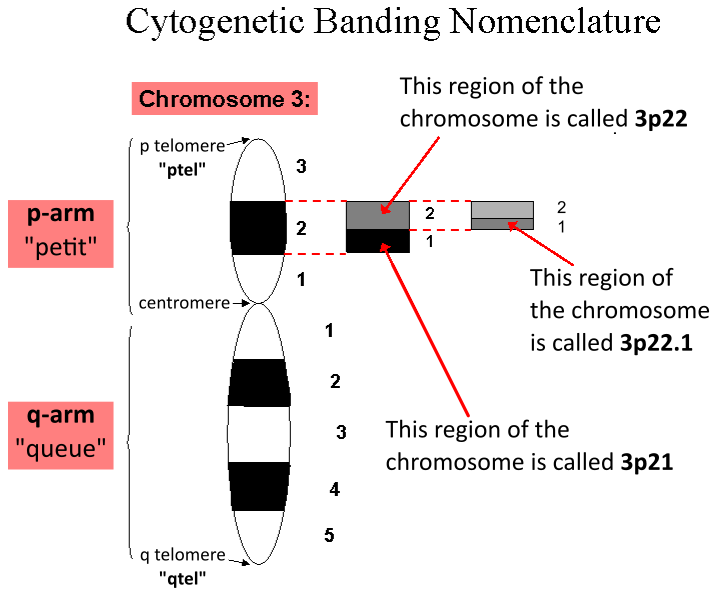
Locus (genetics)
In genetics, a locus (plural loci) is a specific, fixed position on a chromosome where a particular gene or genetic marker is located. Each chromosome carries many genes, with each gene occupying a different position or locus; in humans, the total number of protein-coding genes in a complete haploid set of 23 chromosomes is estimated at 19,000–20,000. Genes may possess multiple variants known as alleles, and an allele may also be said to reside at a particular locus. Diploid and polyploid cells whose chromosomes have the same allele at a given locus are called homozygous with respect to that locus, while those that have different alleles at a given locus are called heterozygous. The ordered list of loci known for a particular genome is called a gene map. Gene mapping is the process of determining the specific locus or loci responsible for producing a particular phenotype or biological trait. Association mapping, also known as "linkage disequilibrium mapping", is a method of ma ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Microarray
A microarray is a multiplex lab-on-a-chip. Its purpose is to simultaneously detect the expression of thousands of genes from a sample (e.g. from a tissue). It is a two-dimensional array on a solid substrate—usually a glass slide or silicon thin-film cell—that assays (tests) large amounts of biological material using high-throughput screening miniaturized, multiplexed and parallel processing and detection methods. The concept and methodology of microarrays was first introduced and illustrated in antibody microarrays (also referred to as antibody matrix) by Tse Wen Chang in 1983 in a scientific publication and a series of patents. The "gene chip" industry started to grow significantly after the 1995 ''Science Magazine'' article by the Ron Davis and Pat Brown labs at Stanford University. With the establishment of companies, such as Affymetrix, Agilent, Applied Microarrays, Arrayjet, Illumina, and others, the technology of DNA microarrays has become the most sophisticated and ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |