|
FERM Domain
In molecular biology, the FERM domain (F for 4.1 protein, E for ezrin, R for radixin and M for moesin) is a widespread protein module involved in localising proteins to the plasma membrane. FERM domains are found in a number of cytoskeletal-associated proteins that associate with various proteins at the interface between the plasma membrane and the cytoskeleton. The FERM domain is located at the N terminus in the majority of proteins in which it is found. Structure and function Ezrin, moesin, and radixin are highly related proteins (ERM protein family), but the other proteins in which the FERM domain is found do not share any region of similarity outside of this domain. ERM proteins are made of three domains, the FERM domain, a central helical domain and a C-terminal tail domain, which binds F-actin. The amino-acid sequence of the FERM domain is highly conserved among ERM proteins and is responsible for membrane association by direct binding to the cytoplasmic domain or tail ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
EPB41
Protein 4.1, also known as Beatty's Protein, is a protein associated with the cytoskeleton that in humans is encoded by the ''EPB41'' gene. Protein 4.1 is a major structural element of the erythrocyte membrane skeleton. It plays a key role in regulating membrane physical properties of mechanical stability and deformability by stabilizing spectrin-actin interaction. Protein 4.1 (80 kD) interacts with spectrin and short actin filaments to form the erythrocyte membrane skeleton. Mutations of spectrin and protein 4.1 are associated with elliptocytosis or spherocytosis and anemia of varying severity. Clinical significance Elliptocytosis is a hematologic disorder characterized by elliptically shaped erythrocytes and a variable degree of hemolytic anemia. Inherited as an autosomal dominant, elliptocytosis results from mutation in any one of several genes encoding proteins of the red cell membrane skeleton. The form discussed here is the one found in the 1950s to be linked to Rh bloo ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein Domains
In molecular biology, a protein domain is a region of a protein's polypeptide chain that is self-stabilizing and that folds independently from the rest. Each domain forms a compact folded three-dimensional structure. Many proteins consist of several domains, and a domain may appear in a variety of different proteins. Molecular evolution uses domains as building blocks and these may be recombined in different arrangements to create proteins with different functions. In general, domains vary in length from between about 50 amino acids up to 250 amino acids in length. The shortest domains, such as zinc fingers, are stabilized by metal ions or disulfide bridges. Domains often form functional units, such as the calcium-binding EF hand domain of calmodulin. Because they are independently stable, domains can be "swapped" by genetic engineering between one protein and another to make chimeric proteins. Background The concept of the domain was first proposed in 1973 by Wetlaufer after ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Conserved Sequence
In evolutionary biology, conserved sequences are identical or similar sequences in nucleic acids ( DNA and RNA) or proteins across species ( orthologous sequences), or within a genome ( paralogous sequences), or between donor and receptor taxa ( xenologous sequences). Conservation indicates that a sequence has been maintained by natural selection. A highly conserved sequence is one that has remained relatively unchanged far back up the phylogenetic tree, and hence far back in geological time. Examples of highly conserved sequences include the RNA components of ribosomes present in all domains of life, the homeobox sequences widespread amongst Eukaryotes, and the tmRNA in Bacteria. The study of sequence conservation overlaps with the fields of genomics, proteomics, evolutionary biology, phylogenetics, bioinformatics and mathematics. History The discovery of the role of DNA in heredity, and observations by Frederick Sanger of variation between animal insulins in 1949, prompt ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
PH Domain
In chemistry, pH (), historically denoting "potential of hydrogen" (or "power of hydrogen"), is a scale used to specify the acidity or basicity of an aqueous solution. Acidic solutions (solutions with higher concentrations of ions) are measured to have lower pH values than basic or alkaline solutions. The pH scale is logarithmic and inversely indicates the concentration of hydrogen ions in the solution.Bates, Roger G. ''Determination of pH: theory and practice''. Wiley, 1973. :\ce = - \log(a_\ce) = -\log( ce\ce M) where M = mol dm−3. At 25 °C (77 °F), solutions with a pH less than 7 are acidic, and solutions with a pH greater than 7 are basic. Solutions with a pH of 7 at this temperature are neutral (i.e. have the same concentration of H+ ions as OH− ions, i.e. pure water). The neutral value of the pH depends on the temperaturebeing lower than 7 if the temperature increases above 25 °C. The pH value can be less than 0 for very concentrated strong acids, ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ubiquitin
Ubiquitin is a small (8.6 kDa) regulatory protein found in most tissues of eukaryotic organisms, i.e., it is found ''ubiquitously''. It was discovered in 1975 by Gideon Goldstein and further characterized throughout the late 1970s and 1980s. Four genes in the human genome code for ubiquitin: UBB, UBC, UBA52 and RPS27A. The addition of ubiquitin to a substrate protein is called ubiquitylation (or, alternatively, ubiquitination or ubiquitinylation). Ubiquitylation affects proteins in many ways: it can mark them for degradation via the proteasome, alter their cellular location, affect their activity, and promote or prevent protein interactions. Ubiquitylation involves three main steps: activation, conjugation, and ligation, performed by ubiquitin-activating enzymes (E1s), ubiquitin-conjugating enzymes (E2s), and ubiquitin ligases (E3s), respectively. The result of this sequential cascade is to bind ubiquitin to lysine residues on the protein substrate via an isopeptide bond, ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Secondary Structure
Protein secondary structure is the three dimensional conformational isomerism, form of ''local segments'' of proteins. The two most common Protein structure#Secondary structure, secondary structural elements are alpha helix, alpha helices and beta sheets, though beta turns and omega loops occur as well. Secondary structure elements typically spontaneously form as an intermediate before the protein protein folding, folds into its three dimensional protein tertiary structure, tertiary structure. Secondary structure is formally defined by the pattern of hydrogen bonds between the Amine, amino hydrogen and carboxyl oxygen atoms in the peptide backbone chain, backbone. Secondary structure may alternatively be defined based on the regular pattern of backbone Dihedral angle#Dihedral angles of proteins, dihedral angles in a particular region of the Ramachandran plot regardless of whether it has the correct hydrogen bonds. The concept of secondary structure was first introduced by Kaj Ulrik ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Crystal Structure
In crystallography, crystal structure is a description of the ordered arrangement of atoms, ions or molecules in a crystal, crystalline material. Ordered structures occur from the intrinsic nature of the constituent particles to form symmetric patterns that repeat along the principal directions of Three-dimensional space (mathematics), three-dimensional space in matter. The smallest group of particles in the material that constitutes this repeating pattern is the unit cell of the structure. The unit cell completely reflects the symmetry and structure of the entire crystal, which is built up by repetitive Translation (geometry), translation of the unit cell along its principal axes. The translation vectors define the nodes of the Bravais lattice. The lengths of the principal axes, or edges, of the unit cell and the angles between them are the lattice constants, also called ''lattice parameters'' or ''cell parameters''. The symmetry properties of the crystal are described by the con ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
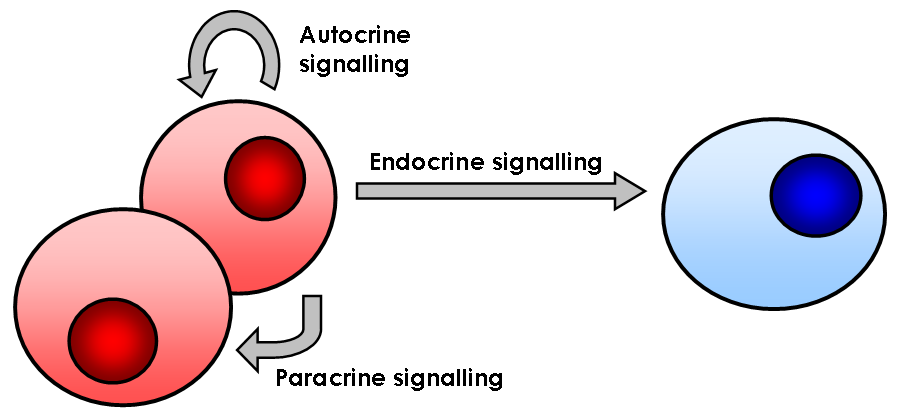
Signalling Pathway
In biology, cell signaling (cell signalling in British English) or cell communication is the ability of a cell to receive, process, and transmit signals with its environment and with itself. Cell signaling is a fundamental property of all cellular life in prokaryotes and eukaryotes. Signals that originate from outside a cell (or extracellular signals) can be physical agents like mechanical pressure, voltage, temperature, light, or chemical signals (e.g., small molecules, peptides, or gas). Cell signaling can occur over short or long distances, and as a result can be classified as autocrine, juxtacrine, intracrine, paracrine, or endocrine. Signaling molecules can be synthesized from various biosynthetic pathways and released through passive or active transports, or even from cell damage. Receptors play a key role in cell signaling as they are able to detect chemical signals or physical stimuli. Receptors are generally proteins located on the cell surface or within the interior ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
GTPase
GTPases are a large family of hydrolase enzymes that bind to the nucleotide guanosine triphosphate (GTP) and hydrolyze it to guanosine diphosphate (GDP). The GTP binding and hydrolysis takes place in the highly conserved P-loop "G domain", a protein domain common to many GTPases. Functions GTPases function as molecular switches or timers in many fundamental cellular processes. Examples of these roles include: * Signal transduction in response to activation of cell surface receptors, including transmembrane receptors such as those mediating taste, smell and vision. * Protein biosynthesis (a.k.a. translation) at the ribosome. * Regulation of cell differentiation, proliferation, division and movement. * Translocation of proteins through membranes. * Transport of vesicles within the cell, and vesicle-mediated secretion and uptake, through GTPase control of vesicle coat assembly. GTPases are active when bound to GTP and inactive when bound to GDP. In the generalized recepto ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Nucleotide
Nucleotides are organic molecules consisting of a nucleoside and a phosphate. They serve as monomeric units of the nucleic acid polymers – deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), both of which are essential biomolecules within all life-forms on Earth. Nucleotides are obtained in the diet and are also synthesized from common nutrients by the liver. Nucleotides are composed of three subunit molecules: a nucleobase, a five-carbon sugar (ribose or deoxyribose), and a phosphate group consisting of one to three phosphates. The four nucleobases in DNA are guanine, adenine, cytosine and thymine; in RNA, uracil is used in place of thymine. Nucleotides also play a central role in metabolism at a fundamental, cellular level. They provide chemical energy—in the form of the nucleoside triphosphates, adenosine triphosphate (ATP), guanosine triphosphate (GTP), cytidine triphosphate (CTP) and uridine triphosphate (UTP)—throughout the cell for the many cellular func ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Guanine
Guanine () ( symbol G or Gua) is one of the four main nucleobases found in the nucleic acids DNA and RNA, the others being adenine, cytosine, and thymine (uracil in RNA). In DNA, guanine is paired with cytosine. The guanine nucleoside is called guanosine. With the formula C5H5N5O, guanine is a derivative of purine, consisting of a fused pyrimidine-imidazole ring system with conjugated double bonds. This unsaturated arrangement means the bicyclic molecule is planar. Properties Guanine, along with adenine and cytosine, is present in both DNA and RNA, whereas thymine is usually seen only in DNA, and uracil only in RNA. Guanine has two tautomeric forms, the major keto form (see figures) and rare enol form. It binds to cytosine through three hydrogen bonds. In cytosine, the amino group acts as the hydrogen bond donor and the C-2 carbonyl and the N-3 amine as the hydrogen-bond acceptors. Guanine has the C-6 carbonyl group that acts as the hydrogen bond acceptor, while a group at N ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Actin Filaments
Microfilaments, also called actin filaments, are protein filaments in the cytoplasm of eukaryotic cells that form part of the cytoskeleton. They are primarily composed of polymers of actin, but are modified by and interact with numerous other proteins in the cell. Microfilaments are usually about 7 nm in diameter and made up of two strands of actin. Microfilament functions include cytokinesis, amoeboid movement, cell motility, changes in cell shape, endocytosis and exocytosis, cell contractility, and mechanical stability. Microfilaments are flexible and relatively strong, resisting buckling by multi-piconewton compressive forces and filament fracture by nanonewton tensile forces. In inducing cell motility, one end of the actin filament elongates while the other end contracts, presumably by myosin II molecular motors. Additionally, they function as part of actomyosin-driven contractile molecular motors, wherein the thin filaments serve as tensile platforms for myosin's ATP-dep ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |