|
E3 Ubiquitin Ligase
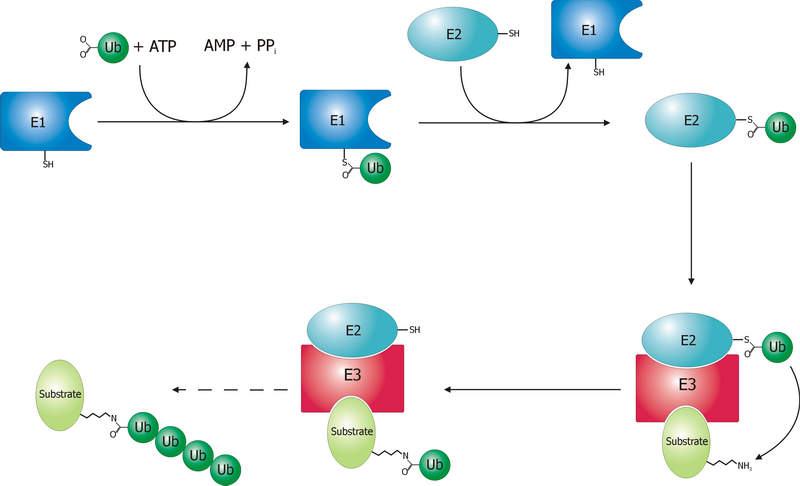
A ubiquitin ligase (also called an E3 ubiquitin ligase) is a protein that recruits an E2 ubiquitin-conjugating enzyme that has been loaded with ubiquitin, recognizes a protein substrate, and assists or directly catalyzes the transfer of ubiquitin from the E2 to the protein substrate. In simple and more general terms, the ligase enables movement of ubiquitin from a ubiquitin carrier to another thing (the substrate) by some mechanism. The ubiquitin, once it reaches its destination, ends up being attached by an isopeptide bond to a lysine residue, which is part of the target protein. E3 ligases interact with both the target protein and the E2 enzyme, and so impart substrate specificity to the E2. Commonly, E3s polyubiquitinate their substrate with Lys48-linked chains of ubiquitin, targeting the substrate for destruction by the proteasome. However, many other types of linkages are possible and alter a protein's activity, interactions, or localization. Ubiquitination by E3 ligases re ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, responding to stimuli, providing structure to cells and organisms, and transporting molecules from one location to another. Proteins differ from one another primarily in their sequence of amino acids, which is dictated by the nucleotide sequence of their genes, and which usually results in protein folding into a specific 3D structure that determines its activity. A linear chain of amino acid residues is called a polypeptide. A protein contains at least one long polypeptide. Short polypeptides, containing less than 20–30 residues, are rarely considered to be proteins and are commonly called peptides. The individual amino acid residues are bonded together by peptide bonds and adjacent amino acid residues. The sequence of amino acid ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RING Finger Domain
In molecular biology, a RING (short for Really Interesting New Gene) finger domain is a protein structural domain of zinc finger type which contains a C3HC4 amino acid motif which binds two zinc cations (seven cysteines and one histidine arranged non-consecutively). This protein domain contains 40 to 60 amino acids. Many proteins containing a RING finger play a key role in the ubiquitination pathway. Zinc fingers Zinc finger (Znf) domains are relatively small protein motifs that bind one or more zinc atoms, and which usually contain multiple finger-like protrusions that make tandem contacts with their target molecule. They bind DNA, RNA, protein and/or lipid substrates. Their binding properties depend on the amino acid sequence of the finger domains and of the linker between fingers, as well as on the higher-order structures and the number of fingers. Znf domains are often found in clusters, where fingers can have different binding specificities. There are many superfamilie ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Amino Acid
Amino acids are organic compounds that contain both amino and carboxylic acid functional groups. Although hundreds of amino acids exist in nature, by far the most important are the alpha-amino acids, which comprise proteins. Only 22 alpha amino acids appear in the genetic code. Amino acids can be classified according to the locations of the core structural functional groups, as Alpha and beta carbon, alpha- , beta- , gamma- or delta- amino acids; other categories relate to Chemical polarity, polarity, ionization, and side chain group type (aliphatic, Open-chain compound, acyclic, aromatic, containing hydroxyl or sulfur, etc.). In the form of proteins, amino acid '' residues'' form the second-largest component (water being the largest) of human muscles and other tissues. Beyond their role as residues in proteins, amino acids participate in a number of processes such as neurotransmitter transport and biosynthesis. It is thought that they played a key role in enabling l ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Degron
A degron is a portion of a protein that is important in regulation of protein degradation rates. Known degrons include short amino acid sequences, structural motifs and exposed amino acids (often Lysine or Arginine) located anywhere in the protein. In fact, some proteins can even contain multiple degrons. Degrons are present in a variety of organisms, from the N-degrons (see N-end Rule) first characterized in yeast to the PEST sequence of mouse ornithine decarboxylase. Degrons have been identified in prokaryotes as well as eukaryotes. While there are many types of different degrons, and a high degree of variability even within these groups, degrons are all similar for their involvement in regulating the rate of a protein's degradation. Much like protein degradation (see proteolysis) mechanisms are categorized by their dependence or lack thereof on Ubiquitin, a small protein involved in proteasomal protein degradation, Degrons may also be referred to as “Ubiquitin-dependent" ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cell (biology)
The cell is the basic structural and functional unit of life forms. Every cell consists of a cytoplasm enclosed within a membrane, and contains many biomolecules such as proteins, DNA and RNA, as well as many small molecules of nutrients and metabolites.Cell Movements and the Shaping of the Vertebrate Body in Chapter 21 of Molecular Biology of the Cell '' fourth edition, edited by Bruce Alberts (2002) published by Garland Science. The Alberts text discusses how the "cellular building blocks" move to shape developing embryos. It is also common to describe small molecules such as ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Substrate (chemistry)
In chemistry, the term substrate is highly context-dependent. Broadly speaking, it can refer either to a chemical species being observed in a chemical reaction, or to a surface on which other chemical reactions or microscopy are performed. In the former sense, a reagent is added to the ''substrate'' to generate a product through a chemical reaction. The term is used in a similar sense in synthetic and organic chemistry, where the substrate is the chemical of interest that is being modified. In biochemistry, an enzyme substrate is the material upon which an enzyme acts. When referring to Le Chatelier's principle, the substrate is the reagent whose concentration is changed. ;Spontaneous reaction : :*Where S is substrate and P is product. ;Catalysed reaction : :*Where S is substrate, P is product and C is catalyst. In the latter sense, it may refer to a surface on which other chemical reactions are performed or play a supporting role in a variety of spectroscopic and micro ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
SH2 Domain
The SH2 (Src Homology 2) domain is a structurally conserved protein domain contained within the Src oncoprotein and in many other intracellular signal-transducing proteins. SH2 domains allow proteins containing those domains to dock to phosphorylated tyrosine residues on other proteins. SH2 domains are commonly found in adaptor proteins that aid in the signal transduction of receptor tyrosine kinase pathways. Background SH2 is conserved by signalization of protein tyrosine kinase, which are binding on phosphotyrosine (pTyr). In the human proteome the class of pTyr-selective recognition domains is represented by SH2 domains. The N-terminal SH2 domains of cytoplasmic tyrosine kinase was at the beginning of evolution evolved with the occurrence of tyrosine phosphorylation. At the beginning it was supposed that, these domains serve as a substrate for their target kinase. Protein-protein interactions play a major role in cellular growth and development. Modular domains, which are t ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Epidermal Growth Factor Receptor
The epidermal growth factor receptor (EGFR; ErbB-1; HER1 in humans) is a transmembrane protein that is a receptor for members of the epidermal growth factor family (EGF family) of extracellular protein ligands. The epidermal growth factor receptor is a member of the ErbB family of receptors, a subfamily of four closely related receptor tyrosine kinases: EGFR (ErbB-1), HER2/neu (ErbB-2), Her 3 (ErbB-3) and Her 4 (ErbB-4). In many cancer types, mutations affecting EGFR expression or activity could result in cancer. Epidermal growth factor and its receptor was discovered by Stanley Cohen of Vanderbilt University. Cohen shared the 1986 Nobel Prize in Medicine with Rita Levi-Montalcini for their discovery of growth factors. Deficient signaling of the EGFR and other receptor tyrosine kinases in humans is associated with diseases such as Alzheimer's, while over-expression is associated with the development of a wide variety of tumors. Interruption of EGFR signalling, either ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Endocytosis
Endocytosis is a cellular process in which substances are brought into the cell. The material to be internalized is surrounded by an area of cell membrane, which then buds off inside the cell to form a vesicle containing the ingested material. Endocytosis includes pinocytosis (cell drinking) and phagocytosis (cell eating). It is a form of active transport. History The term was proposed by De Duve in 1963. Phagocytosis was discovered by Élie Metchnikoff in 1882. Pathways Endocytosis pathways can be subdivided into four categories: namely, receptor-mediated endocytosis (also known as clathrin-mediated endocytosis), caveolae, pinocytosis, and phagocytosis. * Clathrin-mediated endocytosis is mediated by the production of small (approx. 100 nm in diameter) vesicles that have a morphologically characteristic coat made up of the cytosolic protein clathrin. Clathrin-coated vesicles (CCVs) are found in virtually all cells and form domains of the plasma membrane ter ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ubiquitin Lysines
Ubiquitin is a small (8.6 kDa) regulatory protein found in most tissues of eukaryotic organisms, i.e., it is found ''ubiquitously''. It was discovered in 1975 by Gideon Goldstein and further characterized throughout the late 1970s and 1980s. Four genes in the human genome code for ubiquitin: UBB, UBC, UBA52 and RPS27A. The addition of ubiquitin to a substrate protein is called ubiquitylation (or, alternatively, ubiquitination or ubiquitinylation). Ubiquitylation affects proteins in many ways: it can mark them for degradation via the proteasome, alter their cellular location, affect their activity, and promote or prevent protein interactions. Ubiquitylation involves three main steps: activation, conjugation, and ligation, performed by ubiquitin-activating enzymes (E1s), ubiquitin-conjugating enzymes (E2s), and ubiquitin ligases (E3s), respectively. The result of this sequential cascade is to bind ubiquitin to lysine residues on the protein substrate via an isopeptide bond, ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cullin
Cullins are a family of hydrophobic scaffold proteins which provide support for ubiquitin ligases (E3). All eukaryotes appear to have cullins. They combine with RING proteins to form ''Cullin-RING ubiquitin ligases'' (CRLs) that are highly diverse and play a role in myriad cellular processes, most notably protein degradation by ubiquitination. The human genome contains eight cullin genes * CUL1, part of SCF complex * CUL2, part of ECS complex ( Elongin C - CUL2 - SOCS-box) * CUL3, part of CUL3-BTB complex * CUL4A * CUL4B * CUL5 * CUL7 * CUL9, also known as PARC There is also a more distant member called ANAPC2 (or APC2), part of the Anaphase-promoting complex. CUL1, 2, 3, 4A, 4B, 5 and 7 each form part of a multi-subunit ubiquitin complex. Cullin-RING ubiquitin ligases Cullin-RING ubiquitin ligases (CRLs), such as Cul1 (SCF) play an essential role in targeting proteins for ubiquitin-mediated destruction; as such, they are diverse in terms of composition and function, regu ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |