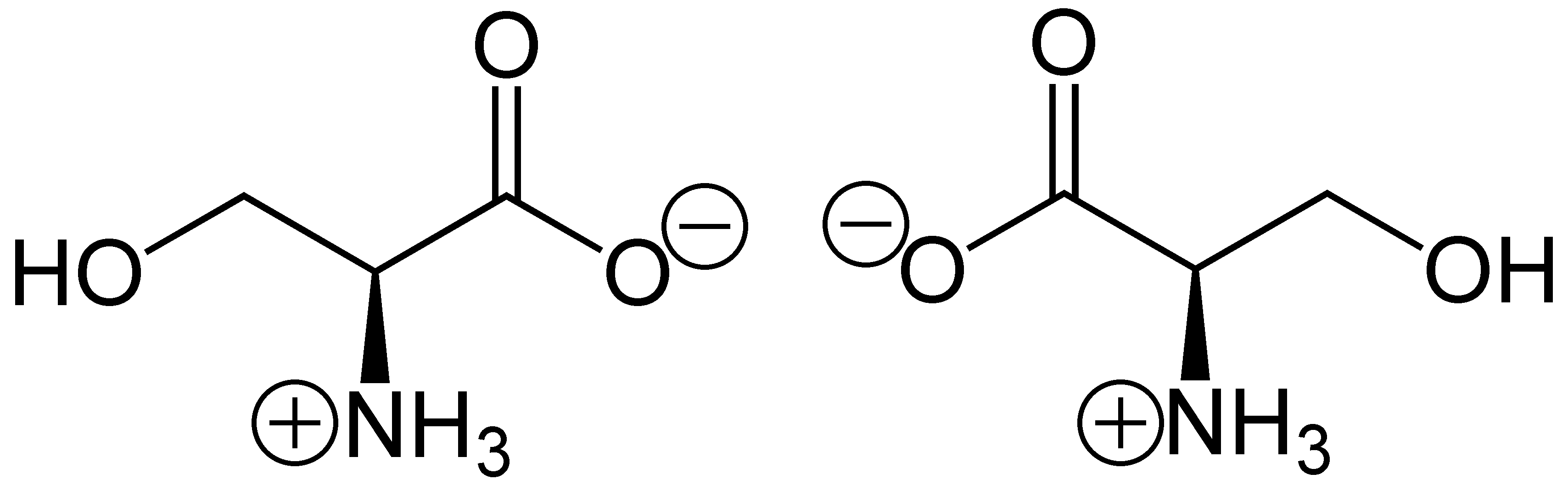|
Deoxyribonuclease IV
Deoxyribonuclease IV (phage-T4-induced) (, ''endodeoxyribonuclease IV (phage T4-induced)'', ''E. coli endonuclease IV'', ''endodeoxyribonuclease'', ''redoxyendonuclease'', ''deoxriboendonuclease'', ''Escherichia coli endonuclease II'', ''endonuclease II'', ''DNA-adenine-transferase'') is catalyzes the degradation nucleotides in DsDNA by attacking the 5'-terminal end. Deoxyribonuclease IV is a type of deoxyribonuclease that has both an exonucleolytic and an endonucleolytic activity. It functions at abasic or apurinic-apyrimidininc sites when the cell is undergoing nucleotide excision repair pathway. In addition, the endonuclease IV consists of several activities such as AP endonuclease, 3'-diesterase, 3'->5' exonuclease, and 3'phosphatase. The endonuclease IV is encoded by denB of bacteriophage T4 and its binding sequence is 5′-dT, , dCdAdCdTdTdC-3′. It has been discovered that serine 176 residue plays a crucial role in increasing the hydrolysis rate of the endonuclease of a c ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Nucleotide Excision Repair
Nucleotide excision repair is a DNA repair mechanism. DNA damage occurs constantly because of chemicals (e.g. intercalating agents), radiation and other mutagens. Three excision repair pathways exist to repair single stranded DNA damage: Nucleotide excision repair (NER), base excision repair (BER), and DNA mismatch repair (MMR). While the BER pathway can recognize specific non-bulky lesions in DNA, it can correct only damaged bases that are removed by specific glycosylases. Similarly, the MMR pathway only targets mismatched Watson-Crick base pairs. Nucleotide excision repair (NER) is a particularly important excision mechanism that removes DNA damage induced by ultraviolet light (UV). UV DNA damage results in bulky DNA adducts - these adducts are mostly thymine dimers and 6,4-photoproducts. Recognition of the damage leads to removal of a short single-stranded DNA segment that contains the lesion. The undamaged single-stranded DNA remains and DNA polymerase uses it as a templa ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Serine
Serine (symbol Ser or S) is an α-amino acid that is used in the biosynthesis of proteins. It contains an α-amino group (which is in the protonated − form under biological conditions), a carboxyl group (which is in the deprotonated − form under biological conditions), and a side chain consisting of a hydroxymethyl group, classifying it as a polar amino acid. It can be synthesized in the human body under normal physiological circumstances, making it a nonessential amino acid. It is encoded by the codons UCU, UCC, UCA, UCG, AGU and AGC. Occurrence This compound is one of the naturally occurring proteinogenic amino acids. Only the L-stereoisomer appears naturally in proteins. It is not essential to the human diet, since it is synthesized in the body from other metabolites, including glycine. Serine was first obtained from silk protein, a particularly rich source, in 1865 by Emil Cramer. Its name is derived from the Latin for silk, ''sericum''. Serine's structure was estab ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Dalton (unit)
The dalton or unified atomic mass unit (symbols: Da or u) is a non-SI unit of mass widely used in physics and chemistry. It is defined as of the mass of an unbound neutral atom of carbon-12 in its nuclear and electronic ground state and at rest. The atomic mass constant, denoted ''m''u, is defined identically, giving . This unit is commonly used in physics and chemistry to express the mass of atomic-scale objects, such as atoms, molecules, and elementary particles, both for discrete instances and multiple types of ensemble averages. For example, an atom of helium-4 has a mass of . This is an intrinsic property of the isotope and all helium-4 atoms have the same mass. Acetylsalicylic acid (aspirin), , has an average mass of approximately . However, there are no acetylsalicylic acid molecules with this mass. The two most common masses of individual acetylsalicylic acid molecules are , having the most common isotopes, and , in which one carbon is carbon-13. The molecular mass ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Exonuclease
Exonucleases are enzymes that work by cleaving nucleotides one at a time from the end (exo) of a polynucleotide chain. A hydrolyzing reaction that breaks phosphodiester bonds at either the 3′ or the 5′ end occurs. Its close relative is the endonuclease, which cleaves phosphodiester bonds in the middle (endo) of a polynucleotide chain. Eukaryotes and prokaryotes have three types of exonucleases involved in the normal turnover of mRNA: 5′ to 3′ exonuclease (Xrn1), which is a dependent decapping protein; 3′ to 5′ exonuclease, an independent protein; and poly(A)-specific 3′ to 5′ exonuclease. In both archaea and eukaryotes, one of the main routes of RNA degradation is performed by the multi-protein exosome complex, which consists largely of 3′ to 5′ exoribonucleases. Significance to polymerase RNA polymerase II is known to be in effect during transcriptional termination; it works with a 5' exonuclease (human gene Xrn2) to degrade the newly formed transcript ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNA Polymerase I
DNA polymerase I (or Pol I) is an enzyme that participates in the process of prokaryotic DNA replication. Discovered by Arthur Kornberg in 1956, it was the first known DNA polymerase (and the first known of any kind of polymerase). It was initially characterized in '' E. coli'' and is ubiquitous in prokaryotes. In ''E. coli'' and many other bacteria, the gene that encodes Pol I is known as ''polA''. The ''E. coli'' Pol I enzyme is composed of 928 amino acids, and is an example of a processive enzyme — it can sequentially catalyze multiple polymerisation steps without releasing the single-stranded template. The physiological function of Pol I is mainly to support repair of damaged DNA, but it also contributes to connecting Okazaki fragments by deleting RNA primers and replacing the ribonucleotides with DNA. Discovery In 1956, Arthur Kornberg and colleagues discovered Pol I by using ''Escherichia coli'' (''E. coli'') extracts to develop a DNA synthesis assay. The scientist ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Escherichia Coli
''Escherichia coli'' (),Wells, J. C. (2000) Longman Pronunciation Dictionary. Harlow ngland Pearson Education Ltd. also known as ''E. coli'' (), is a Gram-negative, facultative anaerobic, rod-shaped, coliform bacterium of the genus ''Escherichia'' that is commonly found in the lower intestine of warm-blooded organisms. Most ''E. coli'' strains are harmless, but some serotypes ( EPEC, ETEC etc.) can cause serious food poisoning in their hosts, and are occasionally responsible for food contamination incidents that prompt product recalls. Most strains do not cause disease in humans and are part of the normal microbiota of the gut; such strains are harmless or even beneficial to humans (although these strains tend to be less studied than the pathogenic ones). For example, some strains of ''E. coli'' benefit their hosts by producing vitamin K2 or by preventing the colonization of the intestine by pathogenic bacteria. These mutually beneficial relationships between ''E. col ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Magnesium
Magnesium is a chemical element with the symbol Mg and atomic number 12. It is a shiny gray metal having a low density, low melting point and high chemical reactivity. Like the other alkaline earth metals (group 2 of the periodic table) it occurs naturally only in combination with other elements and it almost always has an oxidation state of +2. It reacts readily with air to form a thin passivation coating of magnesium oxide that inhibits further corrosion of the metal. The free metal burns with a brilliant-white light. The metal is obtained mainly by electrolysis of magnesium salts obtained from brine. It is less dense than aluminium and is used primarily as a component in strong and lightweight alloys that contain aluminium. In the cosmos, magnesium is produced in large, aging stars by the sequential addition of three helium nuclei to a carbon nucleus. When such stars explode as supernovas, much of the magnesium is expelled into the interstellar medium where it ma ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Mg²⁺
Magnesium is a chemical element with the symbol Mg and atomic number 12. It is a shiny gray metal having a low density, low melting point and high chemical reactivity. Like the other alkaline earth metals (group 2 of the periodic table) it occurs naturally only in combination with other elements and it almost always has an oxidation state of +2. It reacts readily with air to form a thin passivation coating of magnesium oxide that inhibits further corrosion of the metal. The free metal burns with a brilliant-white light. The metal is obtained mainly by electrolysis of magnesium salts obtained from brine. It is less dense than aluminium and is used primarily as a component in strong and lightweight alloys that contain aluminium. In the cosmos, magnesium is produced in large, aging stars by the sequential addition of three helium nuclei to a carbon nucleus. When such stars explode as supernovas, much of the magnesium is expelled into the interstellar medium where it ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Phage T4
Escherichia virus T4 is a species of bacteriophages that infect ''Escherichia coli'' bacteria. It is a double-stranded DNA virus in the subfamily ''Tevenvirinae'' from the family Myoviridae. T4 is capable of undergoing only a lytic lifecycle and not the lysogenic lifecycle. The species was formerly named T-even bacteriophage, a name which also encompasses, among other strains (or isolates), Enterobacteria phage T2, Enterobacteria phage T4 and Enterobacteria phage T6. Use in research Dating back to the 1940s and continuing today, T-even phages are considered the best studied model organisms. Model organisms are usually required to be simple with as few as five genes. Yet, T-even phages are in fact among the largest and highest complexity virus, in which these phage's genetic information is made up of around 300 genes. Coincident with their complexity, T-even viruses were found to have the unusual base hydroxymethylcytosine (HMC) in place of the nucleic acid base cytosine. ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |