|
Cre-Lox Recombination
Cre-Lox recombination is a site-specific recombinase technology, used to carry out deletions, insertions, translocations and inversions at specific sites in the DNA of cells. It allows the DNA modification to be targeted to a specific cell type or be triggered by a specific external stimulus. It is implemented both in eukaryotic and prokaryotic systems. The Cre-lox recombination system has been particularly useful to help neuroscientists to study the brain in which complex cell types and neural circuits come together to generate cognition and behaviors. NIH Blueprint for Neuroscience Research has created several hundreds of Cre driver mouse lines which are currently used by the worldwide neuroscience community. The system consists of a single enzyme, Cre recombinase, that recombines a pair of short target sequences called the ''Lox'' sequences. This system can be implemented without inserting any extra supporting proteins or sequences. The Cre enzyme and the original ''Lox'' ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Site-specific Recombinase Technology
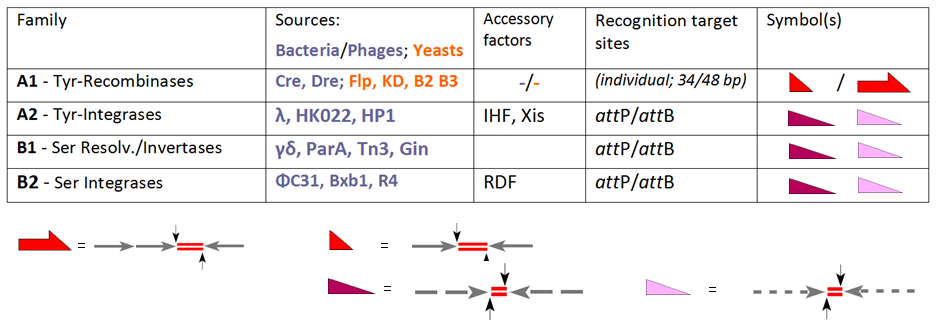
Site-specific recombinase technologies are genome engineering tools that depend on recombinase enzymes to replace targeted sections of DNA. History In the late 1980s gene targeting in murine embryonic stem cells (ESCs) enabled the transmission of mutations into the mouse germ line, and emerged as a novel option to study the genetic basis of regulatory networks as they exist in the genome. Still, classical gene targeting proved to be limited in several ways as gene functions became irreversibly destroyed by the marker gene that had to be introduced for selecting recombinant ESCs. These early steps led to animals in which the mutation was present in all cells of the body from the beginning leading to complex phenotypes and/or early lethality. There was a clear need for methods to restrict these mutations to specific points in development and specific cell types. This dream became reality when groups in the USA were able to introduce bacteriophage and yeast-derived site-specific re ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Dimer (chemistry)
A dimer () (''di-'', "two" + ''-mer'', "parts") is an oligomer consisting of two monomers joined by bonds that can be either strong or weak, covalent or intermolecular. Dimers also have significant implications in polymer chemistry, inorganic chemistry, and biochemistry. The term ''homodimer'' is used when the two molecules are identical (e.g. A–A) and ''heterodimer'' when they are not (e.g. A–B). The reverse of dimerization is often called dissociation. When two oppositely charged ions associate into dimers, they are referred to as ''Bjerrum pairs'', after Niels Bjerrum. Noncovalent dimers Anhydrous carboxylic acids form dimers by hydrogen bonding of the acidic hydrogen and the carbonyl oxygen. For example, acetic acid forms a dimer in the gas phase, where the monomer units are held together by hydrogen bonds. Under special conditions, most OH-containing molecules form dimers, e.g. the water dimer. Excimers and exciplexes are excited structures with a short life ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Holliday Junction
A Holliday junction is a branched nucleic acid structure that contains four double-stranded arms joined. These arms may adopt one of several conformations depending on buffer salt concentrations and the sequence of nucleobases closest to the junction. The structure is named after Robin Holliday, the molecular biologist who proposed its existence in 1964. In biology, Holliday junctions are a key intermediate in many types of genetic recombination, as well as in double-strand break repair. These junctions usually have a symmetrical sequence and are thus mobile, meaning that the four individual arms may slide through the junction in a specific pattern that largely preserves base pairing. Additionally, four-arm junctions similar to Holliday junctions appear in some functional RNA molecules. Immobile Holliday junctions, with asymmetrical sequences that lock the strands in a specific position, were artificially created by scientists to study their structure as a model for natu ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Recombinase-mediated Cassette Exchange
RMCE (recombinase-mediated cassette exchange) is a procedure in reverse genetics allowing the systematic, repeated modification of higher eukaryotic genomes by targeted integration, based on the features of site-specific recombination processes (SSRs). For RMCE, this is achieved by the clean exchange of a preexisting gene cassette for an analogous cassette carrying the "gene of interest" (GOI). The genetic modification of mammalian cells is a standard procedure for the production of correctly modified proteins with pharmaceutical relevance. To be successful, the transfer and expression of the transgene has to be highly efficient and should have a largely predictable outcome. Current developments in the field of gene therapy are based on the same principles. Traditional procedures used for transfer of GOIs are not sufficiently reliable, mostly because the relevant epigenetic influences have not been sufficiently explored: transgenes integrate into chromosomes with low efficiency and ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Base Pairs
A base pair (bp) is a fundamental unit of double-stranded nucleic acids consisting of two nucleobases bound to each other by hydrogen bonds. They form the building blocks of the DNA double helix and contribute to the folded structure of both DNA and RNA. Dictated by specific hydrogen bonding patterns, "Watson–Crick" (or "Watson–Crick–Franklin") base pairs ( guanine– cytosine and adenine– thymine) allow the DNA helix to maintain a regular helical structure that is subtly dependent on its nucleotide sequence. The complementary nature of this based-paired structure provides a redundant copy of the genetic information encoded within each strand of DNA. The regular structure and data redundancy provided by the DNA double helix make DNA well suited to the storage of genetic information, while base-pairing between DNA and incoming nucleotides provides the mechanism through which DNA polymerase replicates DNA and RNA polymerase transcribes DNA into RNA. Many DNA-binding p ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Catalytic Site
In biology and biochemistry, the active site is the region of an enzyme where substrate molecules bind and undergo a chemical reaction. The active site consists of amino acid residues that form temporary bonds with the substrate (binding site) and residues that catalyse a reaction of that substrate (catalytic site). Although the active site occupies only ~10–20% of the volume of an enzyme, it is the most important part as it directly catalyzes the chemical reaction. It usually consists of three to four amino acids, while other amino acids within the protein are required to maintain the tertiary structure of the enzymes. Each active site is evolved to be optimised to bind a particular substrate and catalyse a particular reaction, resulting in high specificity. This specificity is determined by the arrangement of amino acids within the active site and the structure of the substrates. Sometimes enzymes also need to bind with some cofactors to fulfil their function. The active sit ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Lambda Phage
''Enterobacteria phage λ'' (lambda phage, coliphage λ, officially ''Escherichia virus Lambda'') is a bacterial virus, or bacteriophage, that infects the bacterial species ''Escherichia coli'' (''E. coli''). It was discovered by Esther Lederberg in 1950. The wild type of this virus has a temperate life cycle that allows it to either reside within the genome of its host through lysogeny or enter into a lytic phase, during which it kills and lyses the cell to produce offspring. Lambda strains, mutated at specific sites, are unable to lysogenize cells; instead, they grow and enter the lytic cycle after superinfecting an already lysogenized cell. The phage particle consists of a head (also known as a capsid), a tail, and tail fibers (see image of virus below). The head contains the phage's double-strand linear DNA genome. During infection, the phage particle recognizes and binds to its host, ''E. coli'', causing DNA in the head of the phage to be ejected through the tail into t ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Integrase
Retroviral integrase (IN) is an enzyme produced by a retrovirus (such as HIV) that integrates—forms covalent links between—its genetic information into that of the host cell it infects. Retroviral INs are not to be confused with phage integrases ( recombinases) used in biotechnology, such as λ phage integrase, as discussed in site-specific recombination. The macromolecular complex of an IN macromolecule bound to the ends of the viral DNA ends has been referred to as the '' intasome''; IN is a key component in this and the retroviral pre-integration complex. Structure All retroviral IN proteins contain three canonical domains, connected by flexible linkers: ● an N-terminal HH-CC zinc-binding domain (a three-helical bundle stabilized by coordination of a Zn(II) cation) ● a catalytic core domain (RNaseH fold) ● a C-terminal DNA-binding domain ( SH3 fold). Crystal and NMR structures of the individual domains and 2-domain constructs of integr ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Amino Acid
Amino acids are organic compounds that contain both amino and carboxylic acid functional groups. Although hundreds of amino acids exist in nature, by far the most important are the alpha-amino acids, which comprise proteins. Only 22 alpha amino acids appear in the genetic code. Amino acids can be classified according to the locations of the core structural functional groups, as Alpha and beta carbon, alpha- , beta- , gamma- or delta- amino acids; other categories relate to Chemical polarity, polarity, ionization, and side chain group type (aliphatic, Open-chain compound, acyclic, aromatic, containing hydroxyl or sulfur, etc.). In the form of proteins, amino acid ''residues'' form the second-largest component ( water being the largest) of human muscles and other tissues. Beyond their role as residues in proteins, amino acids participate in a number of processes such as neurotransmitter transport and biosynthesis. It is thought that they played a key role in enabling li ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
N-terminal End
The N-terminus (also known as the amino-terminus, NH2-terminus, N-terminal end or amine-terminus) is the start of a protein or polypeptide, referring to the free amine group (-NH2) located at the end of a polypeptide. Within a peptide, the amine group is bonded to the carboxylic group of another amino acid, making it a chain. That leaves a free carboxylic group at one end of the peptide, called the C-terminus, and a free amine group on the other end called the N-terminus. By convention, peptide sequences are written N-terminus to C-terminus, left to right (in LTR writing systems). This correlates the translation direction to the text direction, because when a protein is translated from messenger RNA, it is created from the N-terminus to the C-terminus, as amino acids are added to the carboxyl end of the protein. Chemistry Each amino acid has an amine group and a carboxylic group. Amino acids link to one another by peptide bonds which form through a dehydration reaction tha ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
C-terminal End
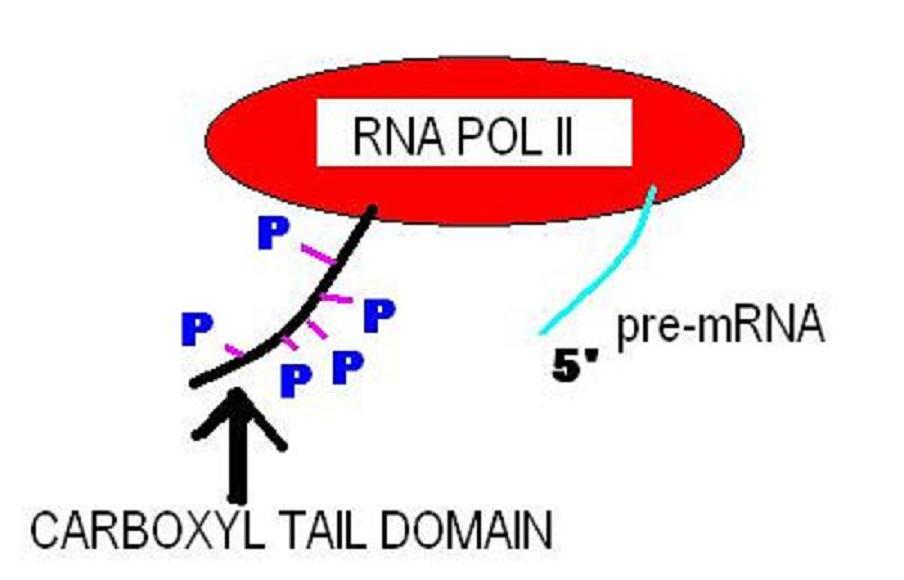
The C-terminus (also known as the carboxyl-terminus, carboxy-terminus, C-terminal tail, C-terminal end, or COOH-terminus) is the end of an amino acid chain (protein or polypeptide), terminated by a free carboxyl group (-COOH). When the protein is translated from messenger RNA, it is created from N-terminus to C-terminus. The convention for writing peptide sequences is to put the C-terminal end on the right and write the sequence from N- to C-terminus. Chemistry Each amino acid has a carboxyl group and an amine group. Amino acids link to one another to form a chain by a dehydration reaction which joins the amine group of one amino acid to the carboxyl group of the next. Thus polypeptide chains have an end with an unbound carboxyl group, the C-terminus, and an end with an unbound amine group, the N-terminus. Proteins are naturally synthesized starting from the N-terminus and ending at the C-terminus. Function C-terminal retention signals While the N-terminus of a protein often con ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |