|
ChIP-chip
ChIP-on-chip (also known as ChIP-chip) is a technology that combines chromatin immunoprecipitation ('ChIP') with DNA microarray (''"chip"''). Like regular ChIP, ChIP-on-chip is used to investigate interactions between proteins and DNA ''in vivo''. Specifically, it allows the identification of the cistrome, the sum of binding sites, for DNA-binding proteins on a genome-wide basis. Whole-genome analysis can be performed to determine the locations of binding sites for almost any protein of interest. As the name of the technique suggests, such proteins are generally those operating in the context of chromatin. The most prominent representatives of this class are transcription factors, replication-related proteins, like origin recognition complex protein (ORC), histones, their variants, and histone modifications. The goal of ChIP-on-chip is to locate protein binding sites that may help identify functional elements in the genome. For example, in the case of a transcription factor as a ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Chromatin Immunoprecipitation
Chromatin immunoprecipitation (ChIP) is a type of immunoprecipitation experimental technique used to investigate the interaction between proteins and DNA in the cell. It aims to determine whether specific proteins are associated with specific genomic regions, such as transcription factors on promoters or other DNA binding sites, and possibly define cistromes. ChIP also aims to determine the specific location in the genome that various histone modifications are associated with, indicating the target of the histone modifiers. ChIP is crucial for the advancements in the field of epigenomics and learning more about epigenetic phenomena. Briefly, the conventional method is as follows: # DNA and associated proteins on chromatin in living cells or tissues are crosslinked (this step is omitted in Native ChIP). # The DNA-protein complexes (chromatin-protein) are then sheared into ~500 bp DNA fragments by sonication or nuclease digestion. # Cross-linked DNA fragments associated with the prot ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
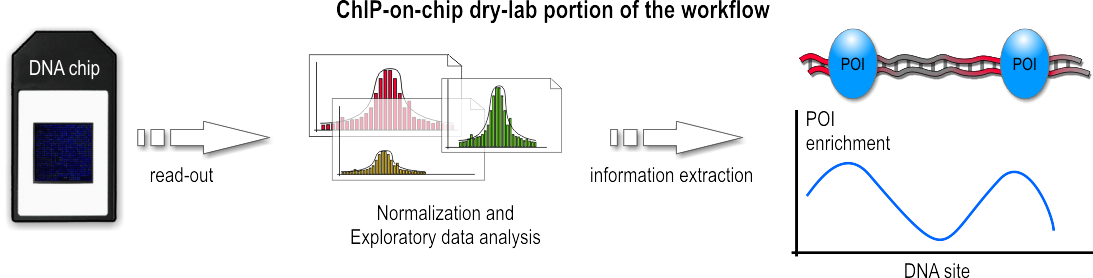
ChIP-on-chip Workflow Overview
ChIP-on-chip (also known as ChIP-chip) is a technology that combines chromatin immunoprecipitation ('ChIP') with DNA microarray (''"chip"''). Like regular ChIP, ChIP-on-chip is used to investigate interactions between proteins and DNA ''in vivo''. Specifically, it allows the identification of the cistrome, the sum of binding sites, for DNA-binding proteins on a genome-wide basis. Whole-genome analysis can be performed to determine the locations of binding sites for almost any protein of interest. As the name of the technique suggests, such proteins are generally those operating in the context of chromatin. The most prominent representatives of this class are transcription factors, replication-related proteins, like origin recognition complex protein (ORC), histones, their variants, and histone modifications. The goal of ChIP-on-chip is to locate protein binding sites that may help identify functional elements in the genome. For example, in the case of a transcription factor as a ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
ChIP-on-chip Wet-lab
ChIP-on-chip (also known as ChIP-chip) is a technology that combines chromatin immunoprecipitation ('ChIP') with DNA microarray (''"chip"''). Like regular ChIP, ChIP-on-chip is used to investigate interactions between proteins and DNA ''in vivo''. Specifically, it allows the identification of the cistrome, the sum of binding sites, for DNA-binding proteins on a genome-wide basis. Whole-genome analysis can be performed to determine the locations of binding sites for almost any protein of interest. As the name of the technique suggests, such proteins are generally those operating in the context of chromatin. The most prominent representatives of this class are transcription factors, replication-related proteins, like origin recognition complex protein (ORC), histones, their variants, and histone modifications. The goal of ChIP-on-chip is to locate protein binding sites that may help identify functional elements in the genome. For example, in the case of a transcription factor as a ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein-DNA Interaction
DNA-binding proteins are proteins that have DNA-binding domains and thus have a specific or general affinity for single- or double-stranded DNA. Sequence-specific DNA-binding proteins generally interact with the major groove of B-DNA, because it exposes more functional groups that identify a base pair. However, there are some known minor groove DNA-binding ligands such as netropsin, distamycin, Hoechst 33258, pentamidine, DAPI and others. Examples DNA-binding proteins include transcription factors which modulate the process of transcription, various polymerases, nucleases which cleave DNA molecules, and histones which are involved in chromosome packaging and transcription in the cell nucleus. DNA-binding proteins can incorporate such domains as the zinc finger, the helix-turn-helix, and the leucine zipper (among many others) that facilitate binding to nucleic acid. There are also more unusual examples such as transcription activator like effectors. Non-specific DNA-protein ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ch-IP
Immunoprecipitation (IP) is the technique of precipitating a protein antigen out of solution using an antibody that specifically binds to that particular protein. This process can be used to isolate and concentrate a particular protein from a sample containing many thousands of different proteins. Immunoprecipitation requires that the antibody be coupled to a solid substrate at some point in the procedure. Types Individual protein immunoprecipitation (IP) Involves using an antibody that is specific for a known protein to isolate that particular protein out of a solution containing many different proteins. These solutions will often be in the form of a crude lysate of a plant or animal tissue. Other sample types could be body fluids or other samples of biological origin. Protein complex immunoprecipitation (Co-IP) Immunoprecipitation of intact protein complexes (i.e. antigen along with any proteins or ligands that are bound to it) is known as co-immunoprecipitation (Co-I ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Gene Regulation
Regulation of gene expression, or gene regulation, includes a wide range of mechanisms that are used by cells to increase or decrease the production of specific gene products (protein or RNA). Sophisticated programs of gene expression are widely observed in biology, for example to trigger developmental pathways, respond to environmental stimuli, or adapt to new food sources. Virtually any step of gene expression can be modulated, from transcriptional initiation, to RNA processing, and to the post-translational modification of a protein. Often, one gene regulator controls another, and so on, in a gene regulatory network. Gene regulation is essential for viruses, prokaryotes and eukaryotes as it increases the versatility and adaptability of an organism by allowing the cell to express protein when needed. Although as early as 1951, Barbara McClintock showed interaction between two genetic loci, Activator (''Ac'') and Dissociator (''Ds''), in the color formation of maize seeds, th ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Chromatin Insulation
An insulator is a type of cis-regulatory element known as a long-range regulatory element. Found in multicellular eukaryotes and working over distances from the promoter element of the target gene, an insulator is typically 300 bp to 2000 bp in length. Insulators contain clustered binding sites for sequence specific DNA-binding proteins and mediate intra- and inter-chromosomal interactions. Insulators function either as an enhancer-blocker or a barrier, or both. The mechanisms by which an insulator performs these two functions include loop formation and nucleosome modifications. There are many examples of insulators, including the CTCF insulator, the ''gypsy'' insulator, and the β-globin locus. The CTCF insulator is especially important in vertebrates, while the ''gypsy'' insulator is implicated in ''Drosophila.'' The β-globin locus was first studied in chicken and then in humans for its insulator activity, both of which utilize CTCF. The genetic implications of insulato ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Affymetrix
Affymetrix is now Applied Biosystems, a brand of DNA microarray products sold by Thermo Fisher Scientific that originated with an American biotechnology research and development and manufacturing company of the same name. The Santa Clara, California-based Affymetrix, Inc. now a part of Thermo Fisher Scientific was co-founded by Alex Zaffaroni and Stephen Fodor. Stephen Fodor and his group, based on their earlier development of methods to fabricate DNA microarrays using semiconductor manufacturing techniques. In 1994, the company's first product under the "GeneChip" Affymetrix trademark, an HIV genotyping chip was introduced, and the company went public in 1996. After incorporation, Affymetrix grew in part by acquiring technologies from other companies, including Genetic MicroSystems (slide-based Microarrays and scanners) and Neomorphic (for bioinformatics) in 2000, ParAllele Bioscience (custom SNP genotyping), USB/Anatrace (biochemical reagents) in 2008, Panomics (low to mid-pl ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Intergenic Region
An intergenic region is a stretch of DNA sequences located between genes. Intergenic regions may contain functional elements and junk DNA. ''Inter''genic regions should not be confused with ''intra''genic regions (or introns), which are non-coding regions that are found ''within'' genes, especially within the genes of eukaryotic organisms. Properties and functions Intergenic regions may contain a number of functional DNA sequences such as promoters and regulatory elements, enhancers, spacers, and (in eukaryotes) centromeres. They may also contain origins of replication, scaffold attachment regions, and transposons and viruses. Non-functional DNA elements such as pseudogenes and repetitive DNA, both of which are types of junk DNA, can also be found in intergenic regions—although they may also be located within genes in introns. As all scientific knowledge is ultimately tentative—and in principle subject to revision given better evidence—it is possible s ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Expression Profiling
In the field of molecular biology, gene expression profiling is the measurement of the activity (the expression) of thousands of genes at once, to create a global picture of cellular function. These profiles can, for example, distinguish between cells that are actively dividing, or show how the cells react to a particular treatment. Many experiments of this sort measure an entire genome simultaneously, that is, every gene present in a particular cell. Several transcriptomics technologies can be used to generate the necessary data to analyse. DNA microarrays measure the relative activity of previously identified target genes. Sequence based techniques, like RNA-Seq, provide information on the sequences of genes in addition to their expression level. Background Expression profiling is a logical next step after sequencing a genome: the sequence tells us what the cell could possibly do, while the expression profile tells us what it is actually doing at a point in time. Genes con ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Open Reading Frame
In molecular biology, open reading frames (ORFs) are defined as spans of DNA sequence between the start and stop codons. Usually, this is considered within a studied region of a prokaryotic DNA sequence, where only one of the six possible reading frames will be "open" (the "reading", however, refers to the RNA produced by transcription of the DNA and its subsequent interaction with the ribosome in translation). Such an ORF may contain a start codon (usually AUG in terms of RNA) and by definition cannot extend beyond a stop codon (usually UAA, UAG or UGA in RNA). That start codon (not necessarily the first) indicates where translation may start. The transcription termination site is located after the ORF, beyond the translation stop codon. If transcription were to cease before the stop codon, an incomplete protein would be made during translation. In eukaryotic genes with multiple exons, introns are removed and exons are then joined together after transcription to yield the final ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
In Situ
''In situ'' (; often not italicized in English) is a Latin phrase that translates literally to "on site" or "in position." It can mean "locally", "on site", "on the premises", or "in place" to describe where an event takes place and is used in many different contexts. For example, in fields such as physics, geology, chemistry, or biology, ''in situ'' may describe the way a measurement is taken, that is, in the same place the phenomenon is occurring without isolating it from other systems or altering the original conditions of the test. The opposite of ''in situ'' is ''ex situ''. Aerospace In the aerospace industry, equipment on-board aircraft must be tested ''in situ'', or in place, to confirm everything functions properly as a system. Individually, each piece may work but interference from nearby equipment may create unanticipated problems. Special test equipment is available for this ''in situ'' testing. It can also refer to repairs made to the aircraft structure or flight con ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |